Team:ETH Zurich/Modeling/Overview
From 2008.igem.org
Luca.Gerosa (Talk | contribs) (→Overview on the modelling framework) |
Luca.Gerosa (Talk | contribs) |
||
| Line 30: | Line 30: | ||
* Is it possible to identify a restriction enzyme that optimizes the probability of reduced genome that retains vital strains?<br><br> | * Is it possible to identify a restriction enzyme that optimizes the probability of reduced genome that retains vital strains?<br><br> | ||
'''Method:'''<br> | '''Method:'''<br> | ||
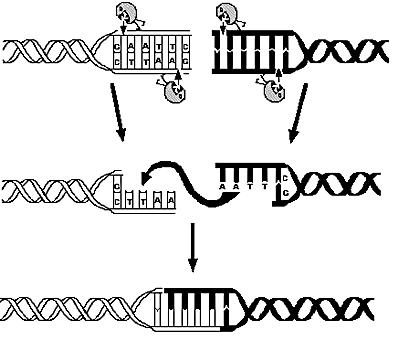
| - | E. | + | E.coli K12 genome was digested using 713 different restriction enzymes and, using annotation information, simple statistical analysis was applied on the calculated fragments. |
<br><br> | <br><br> | ||
'''Results:''' <br> | '''Results:''' <br> | ||
| Line 45: | Line 45: | ||
* Which are the predicted quantitative differences in terms of growth rate and genome size of strains on which has been applied the selection procedure? <br><br> | * Which are the predicted quantitative differences in terms of growth rate and genome size of strains on which has been applied the selection procedure? <br><br> | ||
'''Method:''' <br> | '''Method:''' <br> | ||
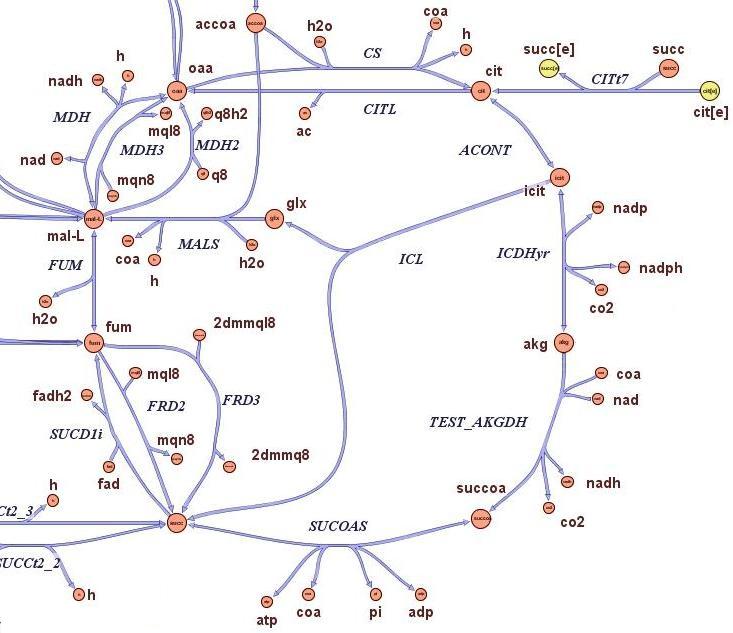
| - | The state-of-the-art genome scale model for E. | + | The state-of-the-art genome scale model for E.coli iAF1260 (1,260 genes included) was modified in order to account for thymidine auxotrophycity, thymidine uptaking limitation, genome reduction and growth on different medium. Stochastic algorithm and flux balance analysis were applied to predict growth rates.<br><br> |
'''Results:''' <br> | '''Results:''' <br> | ||
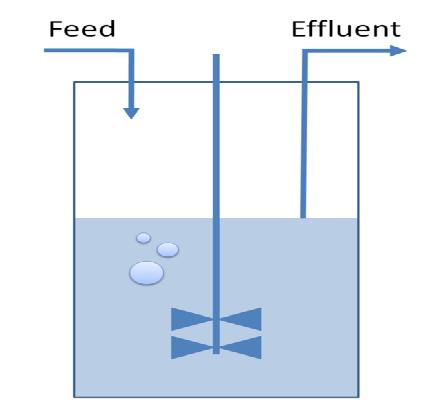
Models show that is indeed possible to select reduced genome strains using thymidine limitation. The quantification shows that the method is at the border line with the sensitivity of chemostat machinery setup for small differencies, but is effective for big reductions (from approximately 10 Kbp on). Predictions show the possibility of reducing up to 61 % of genes for a minimal medium growing strains (corresponding to 59% of chromosome size) and 73 % of genes for rich medium growing strains (corresponding to 71% of chromosome size).<br><br> | Models show that is indeed possible to select reduced genome strains using thymidine limitation. The quantification shows that the method is at the border line with the sensitivity of chemostat machinery setup for small differencies, but is effective for big reductions (from approximately 10 Kbp on). Predictions show the possibility of reducing up to 61 % of genes for a minimal medium growing strains (corresponding to 59% of chromosome size) and 73 % of genes for rich medium growing strains (corresponding to 71% of chromosome size).<br><br> | ||
Revision as of 03:35, 30 October 2008
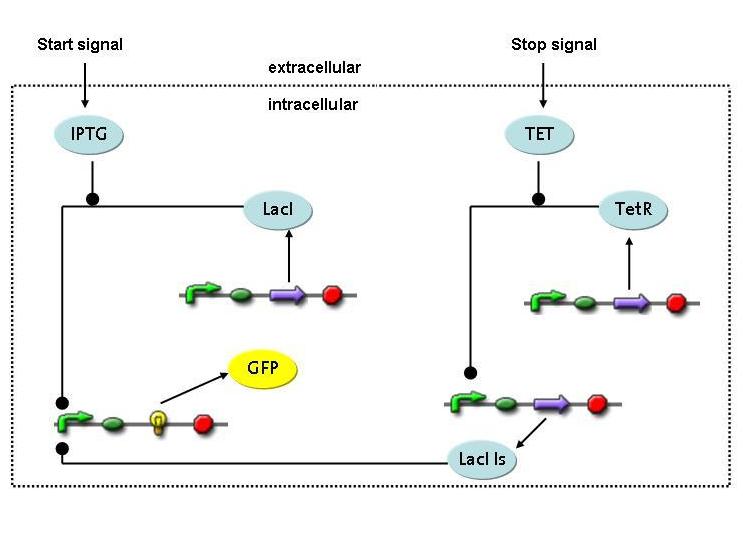
Overview on the modelling frameworkThis page is meant to give an introduction to the overall modelling framework we have constructed in order to assess feasibility analysis, temporal scale details and other parameter estimations with regard of our project setup. As introduced in the Project Overview section, four main components can be identified in the devised mechanism. Accordingly, we divided the modeling framework in four modules that address different challenges.
|
 "
"