Team:BCCS-Bristol/Notebook-Progress
From 2008.igem.org
Progress Reports
- 30th July - 6th August
- BioBrick Transformation
- Heat shock transformations with BioBrick DNA resulted in zero, one or two colonies per attempt
- All colonies have been verified to possess the plasmids
- Electroporation is much more successful
- Agar plug assay
- E. coli MG1655 moved apparently some beads!!!
- but unfortunenately, no chemotaxis was observed
- BioBrick Transformation
- 7th August - 13th August
- BioBrick Transformation
- The self-made electro competent E. coli DH5α cells are working
- A longer incubation time of the punched paper disc in water seems to resolve more BioBrick DNA
- BioBrick Transformation
Progress Report Details
30th July - 6th August
BioBrick Transformation
Last Thursday, the primers [http://partsregistry.org/wiki/index.php/Part:BBa_G00100 VF2] and [http://partsregistry.org/Part:BBa_G00101 VR] arrived. These primers allow a confirmation of a succesful transformation of most BioBrick parts by means of PCR. The resulting fragment length depends on the specific BioBrick DNA.
Using VF2 and VR, a colony PCR are was conducted with all colonies that were obtained with chemical competent cells so far. Two colonies with the [http://partsregistry.org/Part:BBa_J63005 yeast ADH1 promoter] (No. 1 and 2, see left photo) and three colonies with a [http://partsregistry.org/Part:BBa_E0240 GFP generator] (No. 3-5, see left photo) were identified as positive. Thus, all colonies are positive, but the transformation efficiency is too low, since the result was only one or zero colonies per transformation attempt.
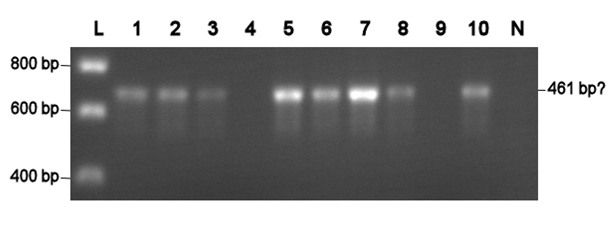
Electroporation should give higher efficiencies. An attempt with a new BioBrick ([http://partsregistry.org/Part:BBa_J63002 ADH1 terminator]) resulted in 69 colonies! Ten of them were analysed using PCR. The resulting fragment should have a length of 461 bp, but from the eigth positive colonies fragments between 650-750 bp were obtained (see right photo). This is probably due to a mutation which might result in a different binding location for one of the primers. Another explanation might be that the denoted length is wrong. Since this BioBrick is not important for our project, it will be disregarded.
|
L= HyperLadder I (BIOLINE) N= Negative control Numbers= See text |
The transformation efficiency of the electroportation was significant higher than of the chemical competent cells. Therefore, an own stock of electric competent cells was made.
Agar plug assay
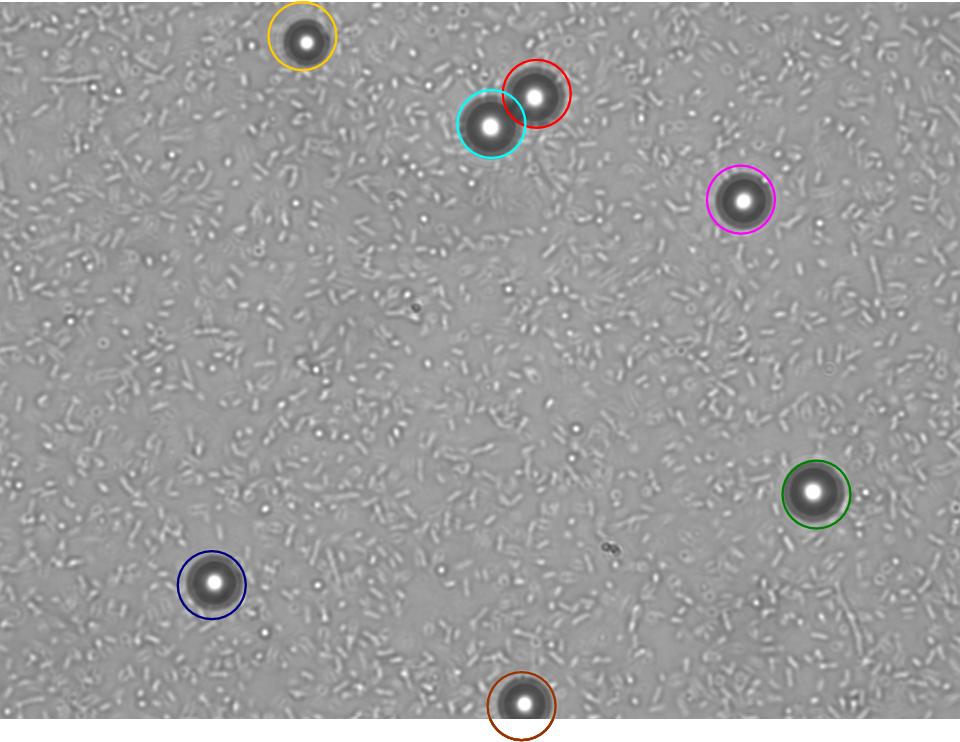
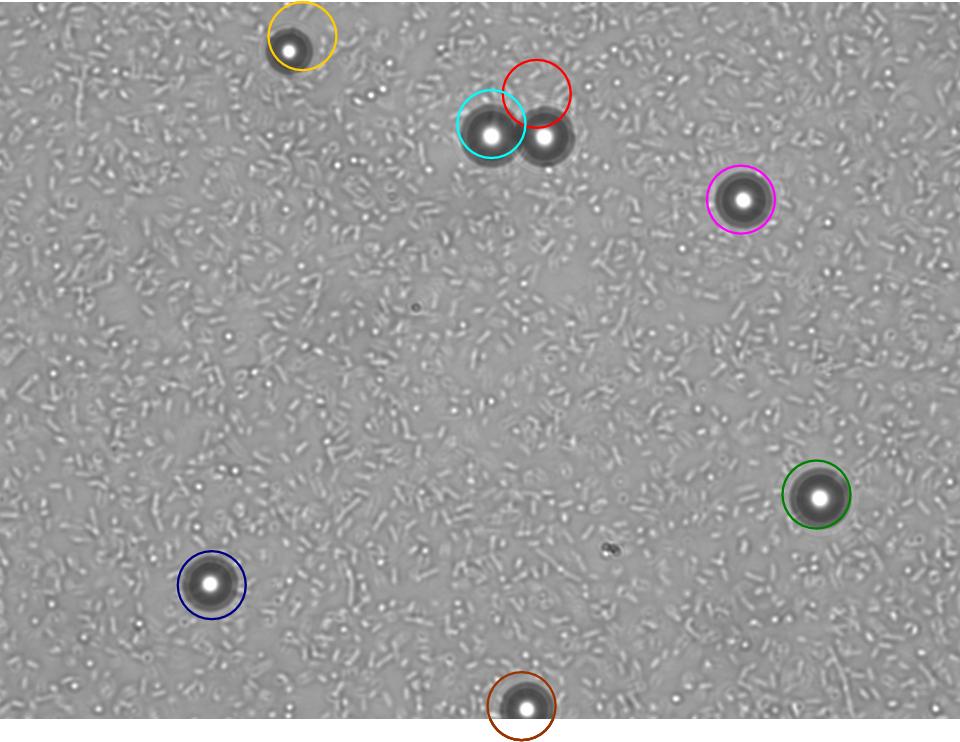
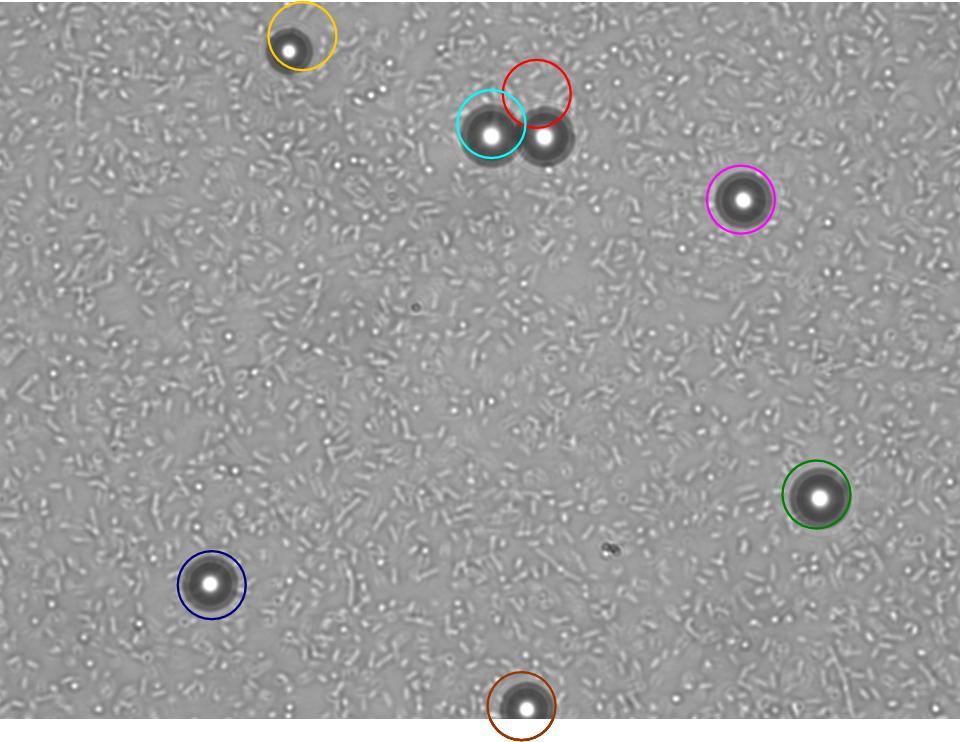
TR235 (E. coli with construct to detect adhesion to hydrophobic surfaces) arrived from Princeton (Thomas J. Silhavy). Cells are viable, growth curve done. 11µm PS beads shown to be moved by bacteria (MG1655); we observed ‘jiggling’ of beads when bacteria present (not seen in no-bacteria control). Also, we observed significant movement in different directions (distances of approx 1x to 5x length of the beads by eye) of beads that were close together, suggesting that movement was not due to currents in the medium. We saw no distance covered in almost all the control beads, a few moved small distances, so some currents are present. Bacteria and beads were left together for ~30mins to allow time for adhesion. No movement has yet been seen when the bacteria and beads are not left to adhere. We will look again at this, as we may have not previously looked closely enough (photographing over a time course of a few minutes).
| 0 mins | 1 min | 2 mins |
|---|
| Circles represent position at 0 mins, one colour for each bead. For example, the bead that starts in red circle moves down and right, while that in the yellow circle moves left. |
|---|
Chemotaxis assay (Yu, H.S. et al. (1997) FEMS Microbiology Letters, 156, 265-269.) has not worked so far. Currents can be seen flowing from the aspartate plug, pushing bacteria away. The focus of this week will be on getting this fixed, with a view to combining chemotaxis and the ability to move beads, allowing us to move beads in a particular direction.
The GRN for switching on the chemotactic response by binding to a bead, and signaling to nearby bacteria that a particle is in the vicinity, is almost finalised.
7th August - 13th August
BioBrick Transformation
The self-made electro competent E. coli DH5α cells were tested twice. The transformation efficiency with pUC19 was 4.45 x 108 cfu/µg and 5.67 x 108 cfu/µg. The first attempt with BioBrick DNA failed, but this might be due to a too low cell amount compared to all the other decanted tubes. In the following transformations, the incubation of the punched paper disc with the DNA was varied in time (3 h and 5.5 h). It seems that a longer incubation increases the transformation efficiency. This has to be confirmed, since the transformations were conducted by different persons. Thereby, next to the [http://partsregistry.org/Part:BBa_J63005 yeast ADH1 promoter] another BioBrick ([http://partsregistry.org/Part:BBa_J63001 enhanced version of EYFP, yeast-optimized YFP]) was tested because the VF2-VR value of [http://partsregistry.org/Part:BBa_J63005 yeast ADH1 promoter] we got didn't coincide with the value given on the iGEM page.
Agar Plug Assay
Evaporation and currents that were occurring in previous attempts at the agarose-in-plug assay were eliminated by ensuring water potentials of plug and chemotaxis buffer were as similar as possible. Both the beads and the bacteria were found to sink to the bottom of the chamber shortly after inoculation. This makes the model much simpler, as everything can be modelled in 2D. We showed that beads need to be left in bacteria for around 30mins in order for significant bead movement to occur. We currently have not determined if this is because the bacteria form a layer under the beads which stops them sticking to the slide, or if adherence itself is needed for bead movement. We determined the diffusion speed of bromophenol blue dye (which is three times larger than aspartate), and concluded that by 30mins aspartate would have diffused throughout the slide.
14th August - 20th August
Agar Plug Assay
We repeated the chemotaxis assays and found that the bacterial density surrounding the plug varied and therefore we obtained more valid results by using lower magnification and scanning around the whole plug (as opposed to focussing on one area for the whole time course). This gave a much better overview of changes in bacterial density.
We tried testing bacterial chemotaxis to 1% serine and 1% aspartate in the plug. Bacterial density around the plug was monitored for 1hr. Bacteria appeared to be repelled by both the serine and aspartate at this concentration compared with the control where no chemoattractant was present in the plug. Research group page (Professor Jong Yoon, Micro/nanofluidic Bio MEMS group) indicates that serine becomes chemorepellent 1M concentration (the same may be true of aspartate).
We have been attempting a 'dead bacteria' control, but have been unable to successfully kill our bacteria without using a method that physically alters them. We have tried sodium azide (bacteriostatic)- but resuspension allows them to recover. Kanamycin and chloramphenicol were also used, but only for a few hours. We might try leaving a culture overnightin these two antibiotics - this should be sufficient to kill them!
 "
"