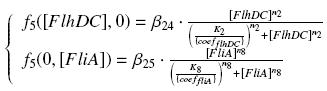|
Method & Algorithm : ƒ5
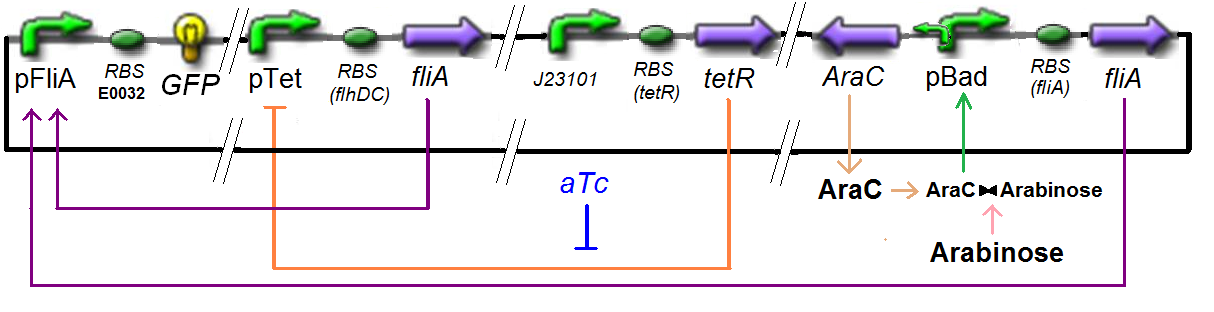 Specific Plasmid Characterisation for ƒ5 We have [FlhDC]real = {coefflhDC}expr(pTet) = {coefflhDC} ƒ1([aTc]i)
and [FliA]real = {coeffliA}expr(pBad) = {coeffliA} ƒ2([arab]i)
but we use [aTc]i = Inv_ƒ1( [FlhDC] )
and [arab]i = Inv_ƒ2( [FliA] )
So, at steady-states,
↓ Table ↑
| param
| signification
| unit
| value
| comments
|
| (fluorescence)
| value of the observed fluorescence
| au
|
| need for 20 mesures with well choosen values of [aTc]i
and for 20 mesures with well choosen values of [arab]i
and 5x5 measures for the relation below?
|
| conversion
| conversion ratio between
fluorescence and concentration
↓ gives ↓
| nM.au-1
| (1/79.429)
|
|
| [GFP]
| GFP concentration at steady-state
| nM
|
|
|
| γGFP
| dilution-degradation rate
of GFP(mut3b)
↓ gives ↓
| min-1
| 0.0198
| Time Cell Division : 35 min.
|
| ƒ5
| activity of
pFliL with RBS E0032
| nM.min-1
|
|
|
| param
| signification
corresponding parameters in the equations
| unit
| value
| comments
|
| β24
| total transcription rate of
FlhDC><pFliL with RBS E0032
β24
| nM.min-1
|
|
|
| (K2/{coeffliA})
| activation constant of FlhDC><pFliL
K2
| nM
|
|
|
| n2
| complexation order of FlhDC><pFliL
n2
| no dimension
|
|
|
| β25
| total transcription rate of
FliA><pFliL with RBS E0032
β25
| nM.min-1
|
|
|
| (K8/{coefflhDC})
| activation constant of FliA><pFliL
K8
| nM
|
|
|
| n8
| complexation order of FliA><pFliL
n8
| no dimension
|
|
|
|
↓ Algorithm ↑
|
find_ƒP
function optimal_parameters = find_FP(X_data, Y_data, initial_parameters)
function output = act_pProm(parameters, X_data)
for k = 1:length(X_data)
output(k) = parameters(1)*hill(X_data(k), parameters(2), parameters(3));
end
end
options=optimset('LevenbergMarquardt','on','TolX',1e-10,'MaxFunEvals',1e10,'TolFun',1e-10,'MaxIter',1e4);
optimal_parameters = lsqcurvefit( @(parameters, X_data) act_pProm(parameters, X_data),...
initial_parameters, X_data, Y_data, options );
end
|
Then, if we have time, we want to verify the expected relation
<Back - to "Implementation" |
<Back - to "Protocol Of Characterization" |
|
 "
"