Team:Heidelberg/Notebook/Sensing Group/Cloning
From 2008.igem.org
(→Fusion receptor) |
|||
| (26 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | __NOTOC__ | ||
<html> | <html> | ||
| Line 466: | Line 467: | ||
==Sensing - Cloning strategy== | ==Sensing - Cloning strategy== | ||
| - | The core part of the sensing project | + | The core part of the sensing project was the construction of the LuxQ-Tar chimeric receptor, which enables the ''E. coli'' killer bacteria to chemotactically respond to a Autoinducer 2 (AI-2) gradient and detect prey cells. The quorum-sensing system was amplified from the ''V. harveyi'' genome while the Tar receptor is from ''E. coli''. On one hand LuxS needed to be cloned and transformed into one cell type, in order to make them produce and secrete AI-2. On the other hand the periplasmic ligand binding domain of LuxQ was fused with the cytoplasmic domain of Tar and cloned on one plasmid together with LuxP which is necessary for AI-2 binding. Generally, restriction sites required for cloning were introduced via the PCR primer. |
| - | ''In silico'' cloning was performed in [http://serialbasics.free.fr/Serial_Cloner.html SerialCloner], Vector maps were designed with PlasMapper [1]. | + | ''In silico'' cloning was performed in [http://serialbasics.free.fr/Serial_Cloner.html SerialCloner], Vector maps were designed with PlasMapper [1]. The sequenecs of used primers can be found in the [[Team:Heidelberg/Notebook/material#Synthetic_oligonucleotides | Materials & Methods]] section. |
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
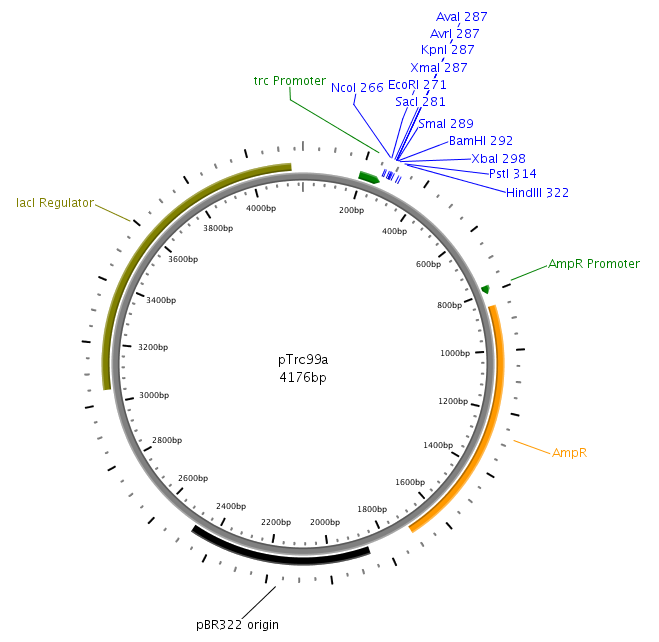
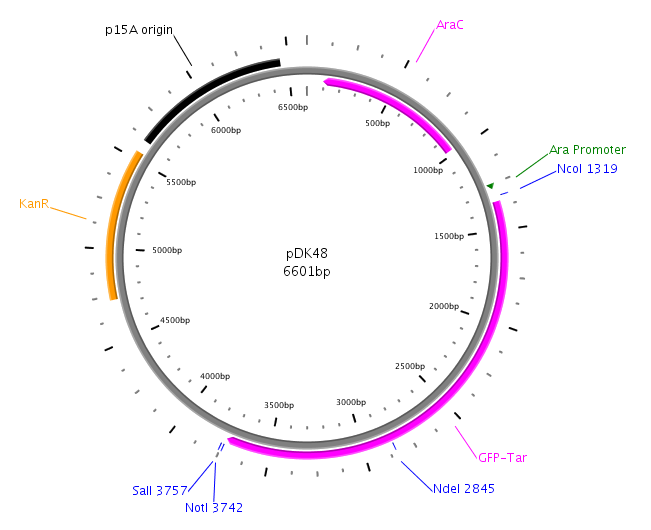
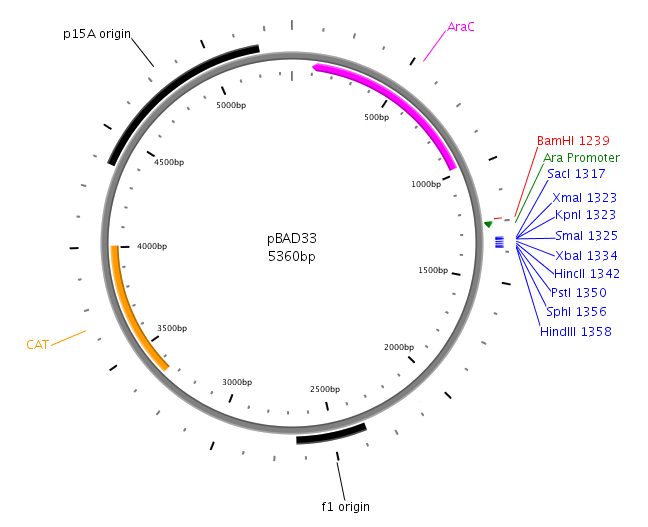
| - | [[Image:HD PTrc99a.png|left|thumb| | + | [[Image:HD PTrc99a.png|left|thumb|230px|pTrc99a for LuxS and LuxQ cloning]][[Image:HD PDK48.png|left|thumb|275px|pDK48 for cloning of fusion]][[Image:HD PBAD33.png|left|thumb|275px|pBAD33 used in Swarm-Assays]] |
<br style="clear: both;"/> | <br style="clear: both;"/> | ||
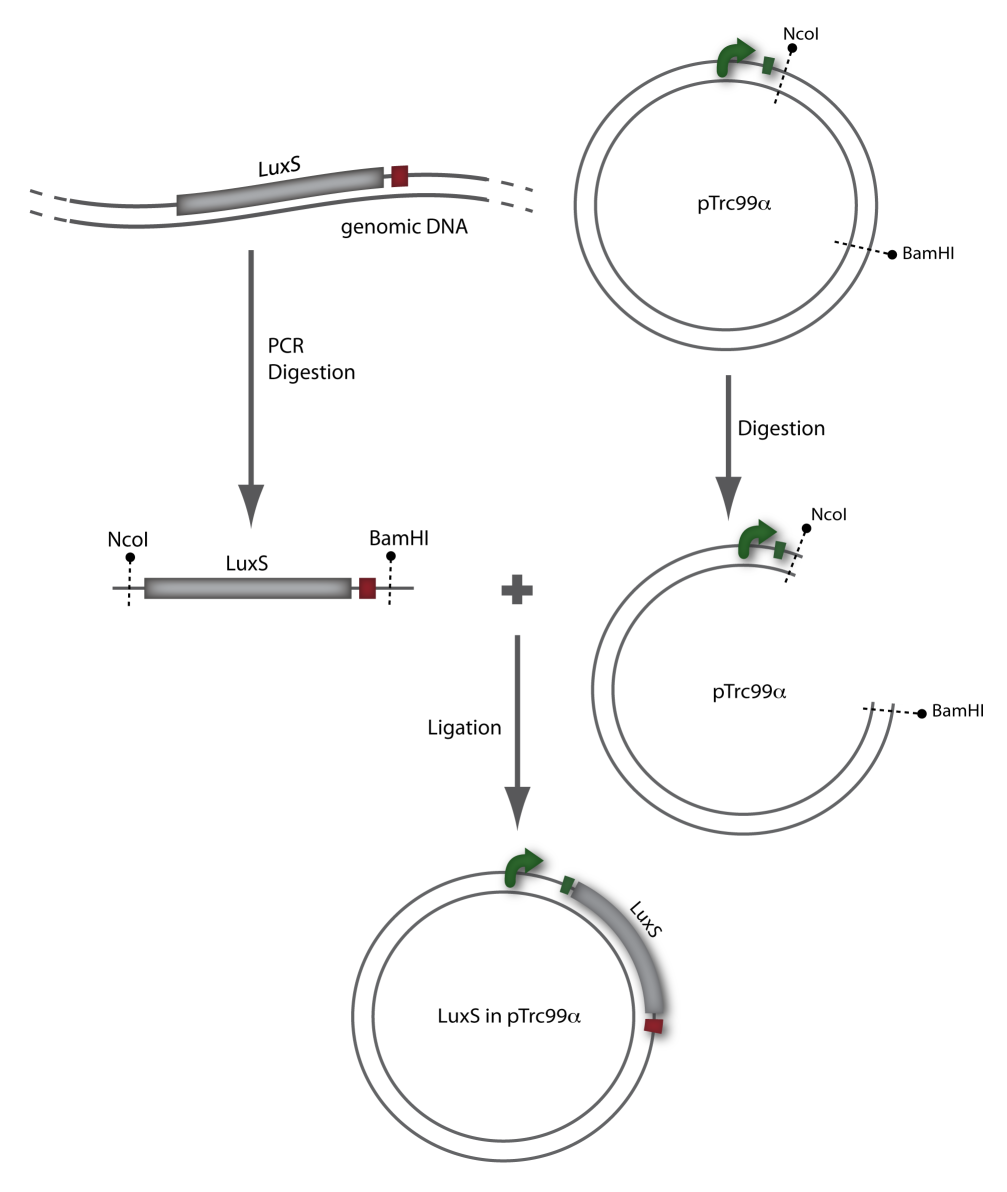
=== LuxS === | === LuxS === | ||
| - | LuxS | + | LuxS was amplified from the ''Vibrio harveyi'' genome using the following primers: LuxSa (forward) and LuxSb (reverse). Subsequently, the product was cloned into the plasmid pTr99alpha using the NcoI and BamHI restriction sites and then transformed into DH5alpha competent cells. |
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
=== LuxQ === | === LuxQ === | ||
| - | LuxQ | + | LuxQ was amplfied with primers LuxQa (forward) and LuxQc (reverse) from the ''V. harveyi'' genome. The obtained fragment was then cloned into the plasmid pTrc99alpha using the NcoI and BamHI restriction sites. Subsequently, the product was used as a template for PCR amplification with respect to cloning of the chimeric receptors (see below). |
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
=== LuxP === | === LuxP === | ||
| - | LuxP | + | LuxP was amplified with primers LuxPc (forward) and LuxPd (reverse) from the ''V. harveyi'' genome. The PCR product was then cloned into the vector pDK48 containing the LuxQ-Tar chimeric receptor sequence. This was done using the restriction sites SalI and NotI. The resulting expression plasmid was used for the transformation of our killer cells. |
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
=== Fusion receptor === | === Fusion receptor === | ||
| - | A detailed description of the Tar receptor and LuxQ quorum-sensing receptor can be found in the [[Team:Heidelberg/Project/Sensing | project description]]. Since it is believed that the linker region (transmembrane domain TM2) is | + | A detailed description of the Tar receptor and LuxQ quorum-sensing receptor can be found in the [[Team:Heidelberg/Project/Sensing | project description]]. Since it is believed that the linker region (transmembrane domain TM2) is critical for signal transduction, two different receptor constructs were designed, referred to as Fusion-1 and Fusion-2. Fusion-1 contained the cytoplasmic domain of Tar (residues 225-564) and the transmembrane domain,TM2 up to the N-terminus of LuxQ (residues 1-298). Fusion-2 contained the cytoplasmic domain as well as the second transmembrane domain, TM2 of Tar (residues 200-564) followed by the periplasmic domain up to the N-terminal end of LuxQ (residues 1-280). |
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
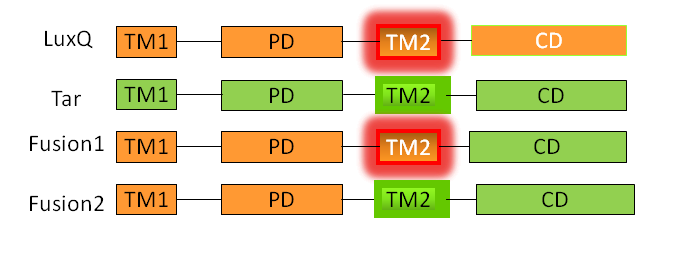
| - | The receptor chimeras | + | [[Image:HD Fusion Schema.png|thumb|center|400px|Schematic structure of the Fusion receptor constructs]] |
| + | |||
| + | The receptor chimeras were generated in two sequential PCR reactions (Fusion PCR). In the first PCR the two single fragments containing parts of the coding sequence of LuxQ (LuxQ-1: primers LuxQa/LuxQ-Tar-1b, LuxQ-2: primers LuxQa/LuxQ-Tar-2b) and Tar (Tar-1: primers LuxQ-Tar-1a/DK9b, Tar-2: primers LuxQ-Tar-2a/DK9b) were amplified. The reverse primer for LuxQ and the forward primer for Tar had complementary ends allowing annealing during the second PCR reaction. In addition to the amplified sequences from the first reaction the forward primer for LuxQ and the reverse primer for Tar were added to the reaction mix after the gel purification in order to be able to obtain the chimeric receptor sequence. Following amplification the chimeric products F1 and F2 were cloned into the expression vector pDK48 via the restriction sites NcoI and NdeI. The presence of the correct sequence was checked by sequencing. | ||
| + | |||
| + | |||
| + | [[Image:HD SensingCloning1.png|thumb|center|500px|Cloning Strategy for LuxS]] | ||
| + | [[Image:HD_SensingCloning2.png|thumb|center|500px|Cloning Strategy for Fusion chimeras]] | ||
=== References === | === References === | ||
[1] Dong, X.; Stothard, P.; Forsythe, I. J. & Wishart, D. S., '''PlasMapper: a web server for drawing and auto-annotating plasmid maps''', Nucleic Acids Res, Vol. 32, pp. W660-W664, 2004 | [1] Dong, X.; Stothard, P.; Forsythe, I. J. & Wishart, D. S., '''PlasMapper: a web server for drawing and auto-annotating plasmid maps''', Nucleic Acids Res, Vol. 32, pp. W660-W664, 2004 | ||
| + | |||
| + | [[https://2008.igem.org/Team:Heidelberg/Notebook/Sensing_Group/Cloning back]] | ||
Latest revision as of 14:12, 29 October 2008


Sensing - Cloning strategy
The core part of the sensing project was the construction of the LuxQ-Tar chimeric receptor, which enables the E. coli killer bacteria to chemotactically respond to a Autoinducer 2 (AI-2) gradient and detect prey cells. The quorum-sensing system was amplified from the V. harveyi genome while the Tar receptor is from E. coli. On one hand LuxS needed to be cloned and transformed into one cell type, in order to make them produce and secrete AI-2. On the other hand the periplasmic ligand binding domain of LuxQ was fused with the cytoplasmic domain of Tar and cloned on one plasmid together with LuxP which is necessary for AI-2 binding. Generally, restriction sites required for cloning were introduced via the PCR primer. In silico cloning was performed in SerialCloner, Vector maps were designed with PlasMapper [1]. The sequenecs of used primers can be found in the Materials & Methods section. [back]
LuxS
LuxS was amplified from the Vibrio harveyi genome using the following primers: LuxSa (forward) and LuxSb (reverse). Subsequently, the product was cloned into the plasmid pTr99alpha using the NcoI and BamHI restriction sites and then transformed into DH5alpha competent cells. [back]
LuxQ
LuxQ was amplfied with primers LuxQa (forward) and LuxQc (reverse) from the V. harveyi genome. The obtained fragment was then cloned into the plasmid pTrc99alpha using the NcoI and BamHI restriction sites. Subsequently, the product was used as a template for PCR amplification with respect to cloning of the chimeric receptors (see below). [back]
LuxP
LuxP was amplified with primers LuxPc (forward) and LuxPd (reverse) from the V. harveyi genome. The PCR product was then cloned into the vector pDK48 containing the LuxQ-Tar chimeric receptor sequence. This was done using the restriction sites SalI and NotI. The resulting expression plasmid was used for the transformation of our killer cells. [back]
Fusion receptor
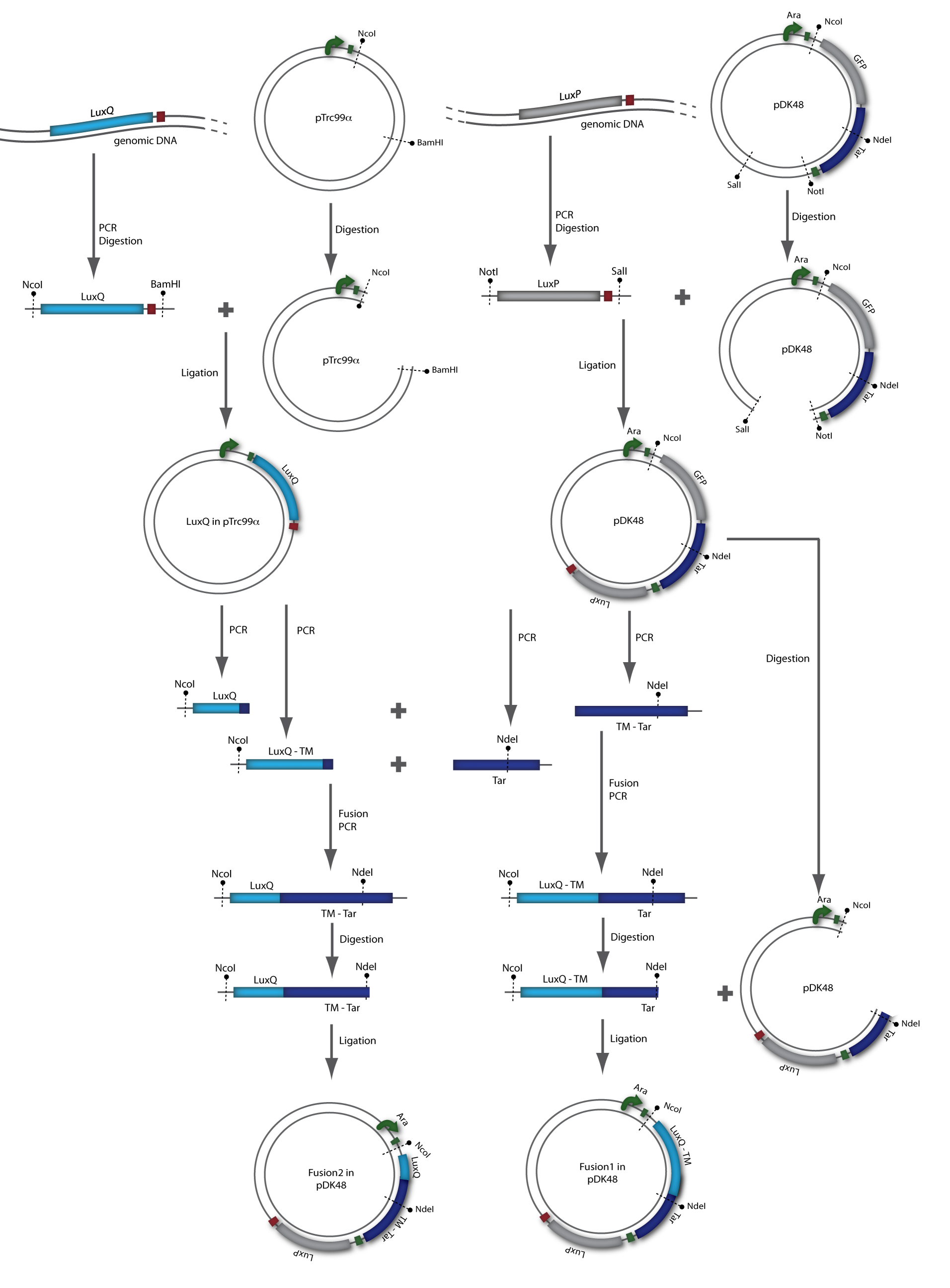
A detailed description of the Tar receptor and LuxQ quorum-sensing receptor can be found in the project description. Since it is believed that the linker region (transmembrane domain TM2) is critical for signal transduction, two different receptor constructs were designed, referred to as Fusion-1 and Fusion-2. Fusion-1 contained the cytoplasmic domain of Tar (residues 225-564) and the transmembrane domain,TM2 up to the N-terminus of LuxQ (residues 1-298). Fusion-2 contained the cytoplasmic domain as well as the second transmembrane domain, TM2 of Tar (residues 200-564) followed by the periplasmic domain up to the N-terminal end of LuxQ (residues 1-280). [back]
The receptor chimeras were generated in two sequential PCR reactions (Fusion PCR). In the first PCR the two single fragments containing parts of the coding sequence of LuxQ (LuxQ-1: primers LuxQa/LuxQ-Tar-1b, LuxQ-2: primers LuxQa/LuxQ-Tar-2b) and Tar (Tar-1: primers LuxQ-Tar-1a/DK9b, Tar-2: primers LuxQ-Tar-2a/DK9b) were amplified. The reverse primer for LuxQ and the forward primer for Tar had complementary ends allowing annealing during the second PCR reaction. In addition to the amplified sequences from the first reaction the forward primer for LuxQ and the reverse primer for Tar were added to the reaction mix after the gel purification in order to be able to obtain the chimeric receptor sequence. Following amplification the chimeric products F1 and F2 were cloned into the expression vector pDK48 via the restriction sites NcoI and NdeI. The presence of the correct sequence was checked by sequencing.
References
[1] Dong, X.; Stothard, P.; Forsythe, I. J. & Wishart, D. S., PlasMapper: a web server for drawing and auto-annotating plasmid maps, Nucleic Acids Res, Vol. 32, pp. W660-W664, 2004
[back]
 "
"