Team:Freiburg/3D-Modeling
From 2008.igem.org
m |
m |
||
| Line 5: | Line 5: | ||
<br><br> | <br><br> | ||
== DNA-Origami == | == DNA-Origami == | ||
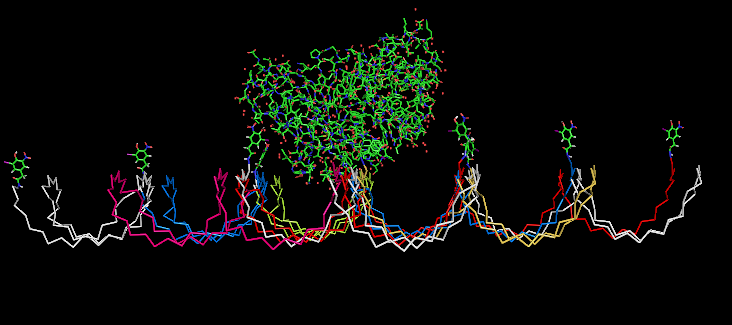
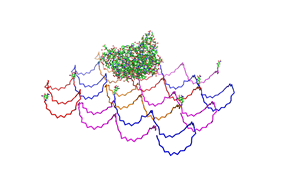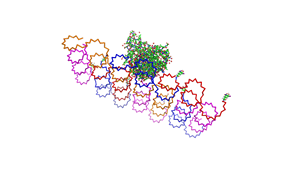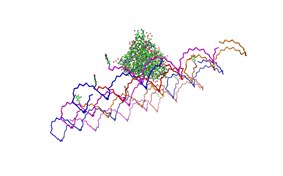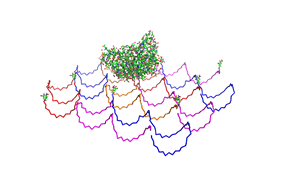
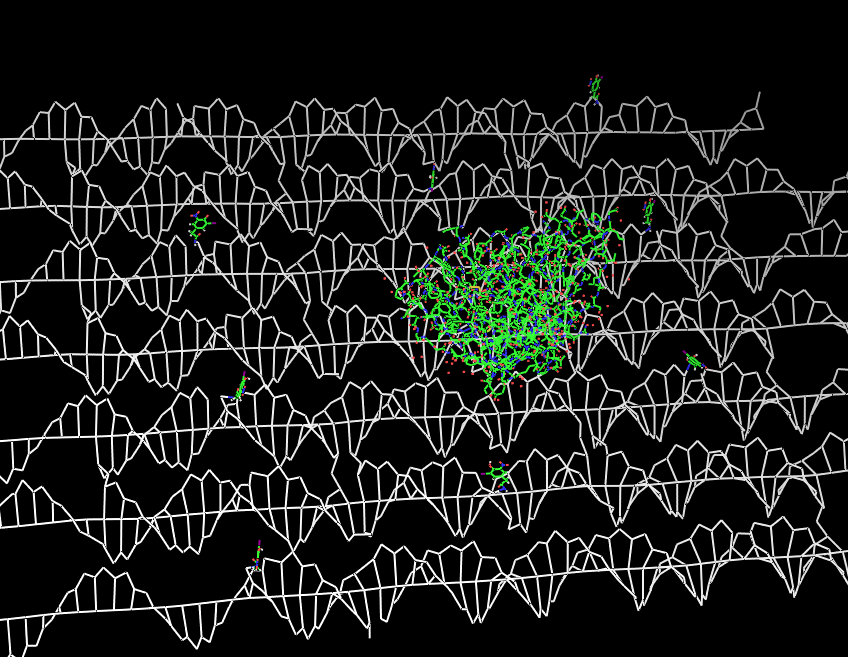
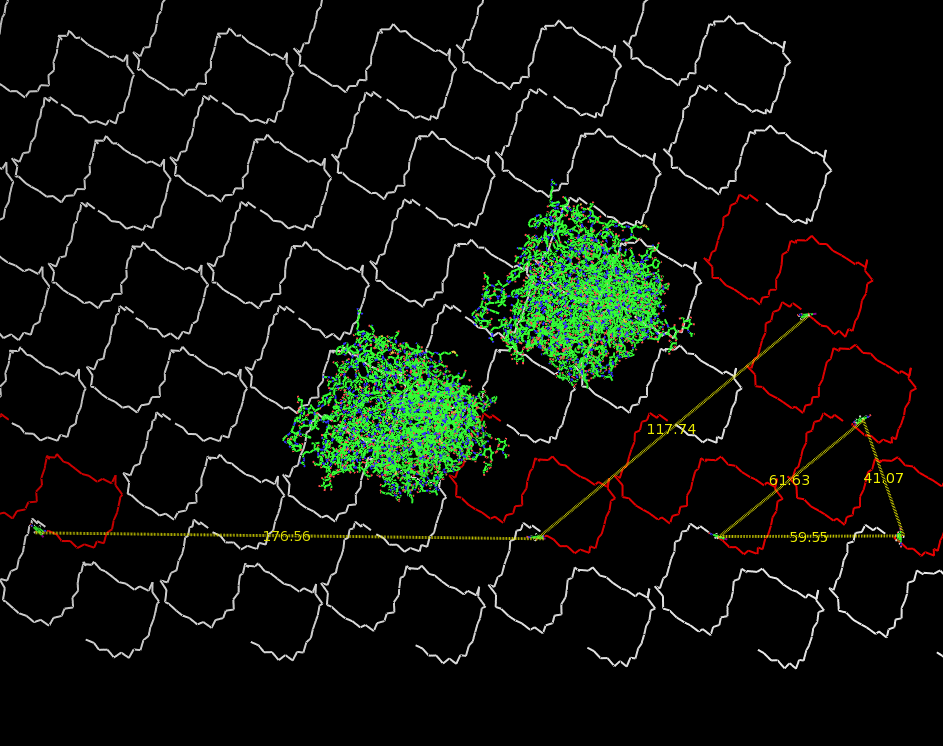
| - | Planning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using [ http://www.nanoengineer-1.com/content/ NanoEngineer], [http://pymol.sourceforge.net/ Pymol] and [http://spdbv.vital-it.ch/ SwissPdbViewer] (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.<br> | + | Planning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using [http://www.nanoengineer-1.com/content/ NanoEngineer], [http://pymol.sourceforge.net/ Pymol] and [http://spdbv.vital-it.ch/ SwissPdbViewer] (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.<br> |
<table> | <table> | ||
<tr> | <tr> | ||
Revision as of 02:11, 30 October 2008
|
_3D-modeling
DNA-OrigamiPlanning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using NanoEngineer, Pymol and SwissPdbViewer (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.
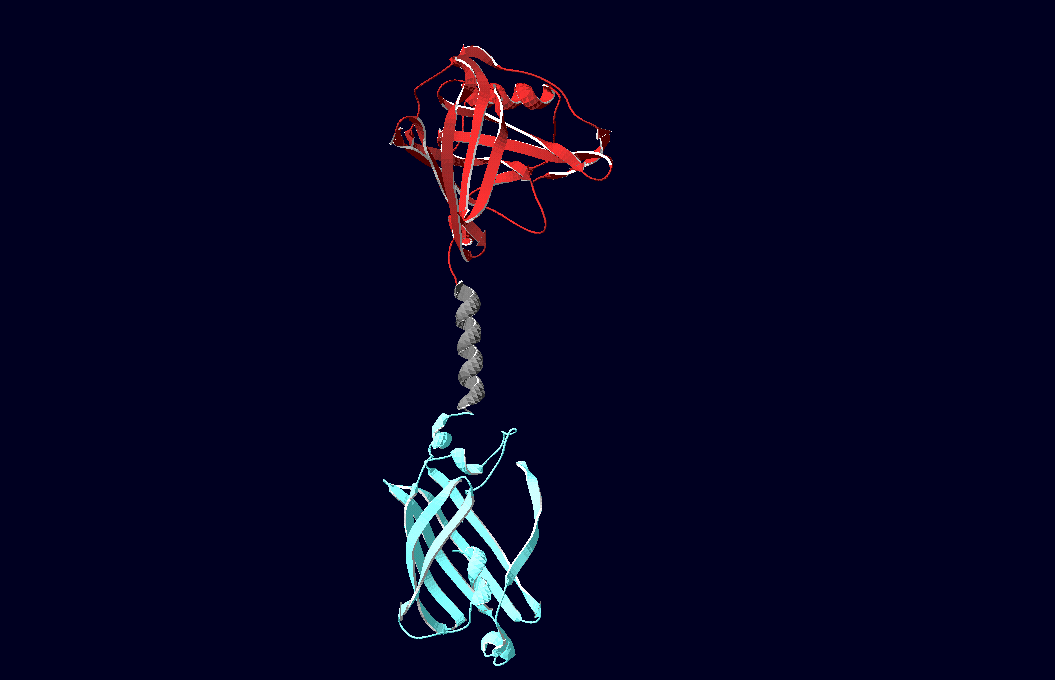
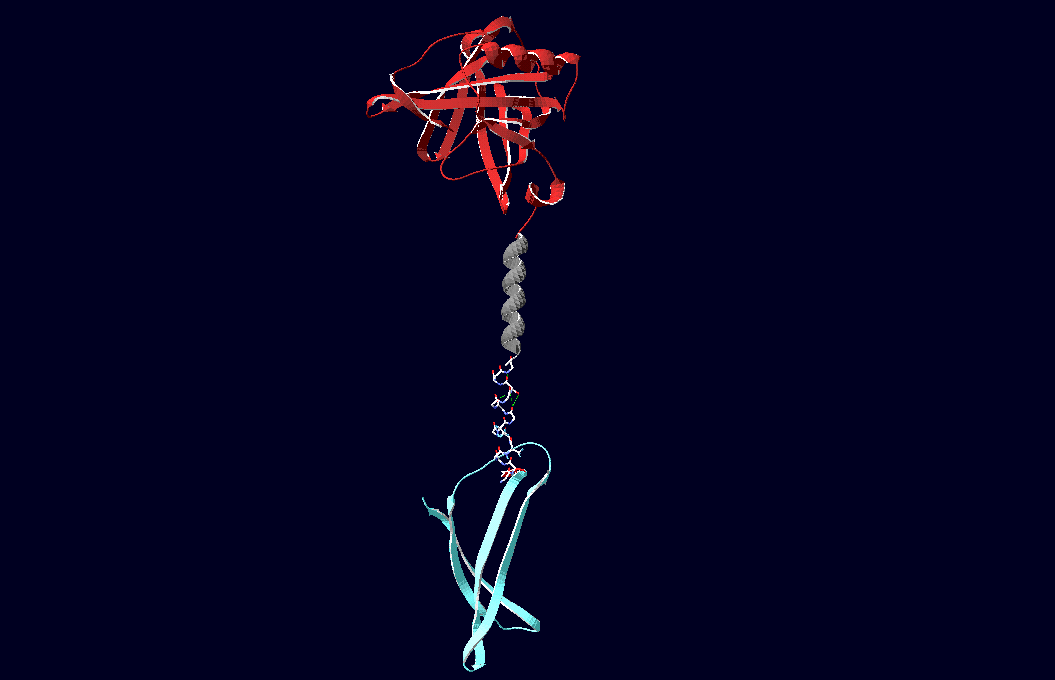
Fusion ProteinsThe pictures below show models of the fusion proteins of our composite parts Bba_K157032 and Bba_K157033 code for (generated with "SwissPdbViewer"): |
 "
"