Team:Freiburg/3D-Modeling
From 2008.igem.org
(Difference between revisions)
m |
|||
| Line 6: | Line 6: | ||
== DNA-Origami == | == DNA-Origami == | ||
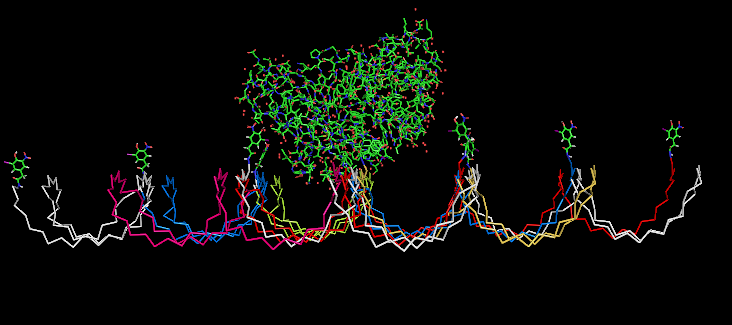
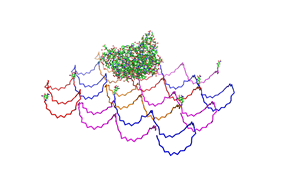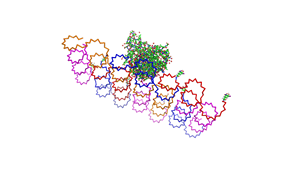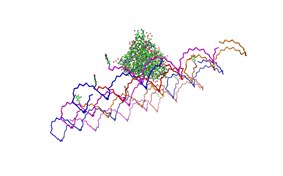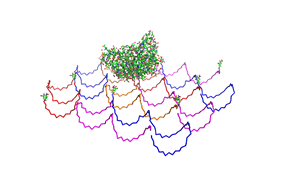
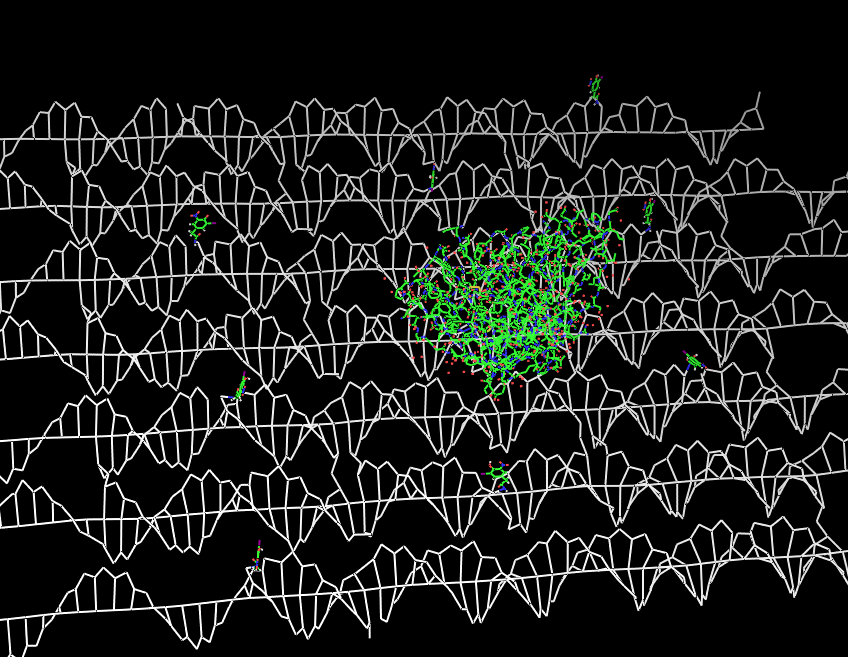
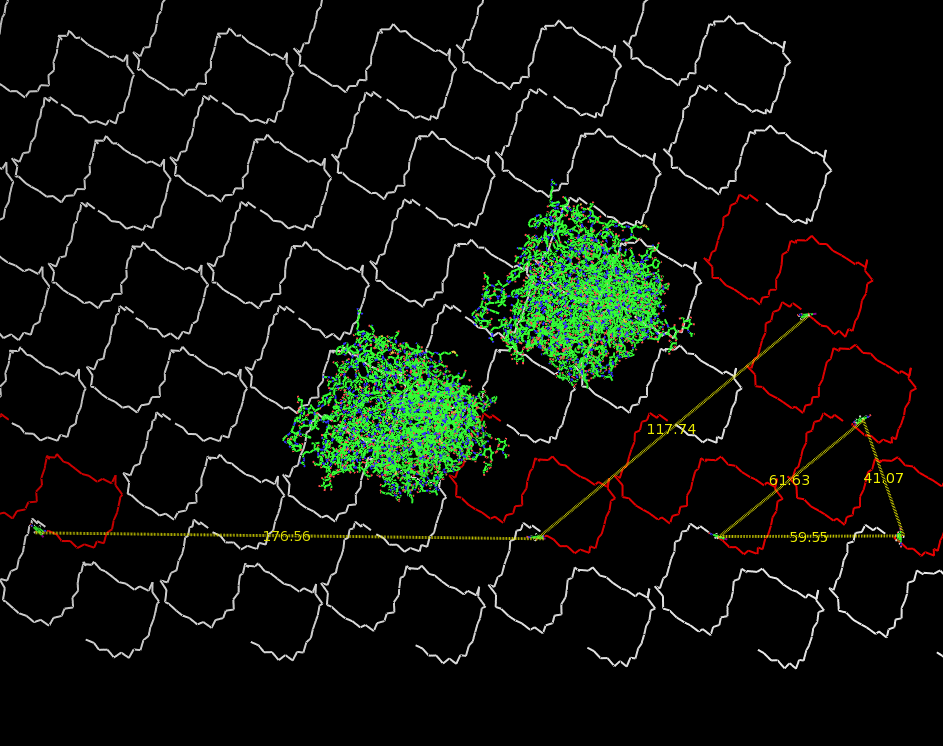
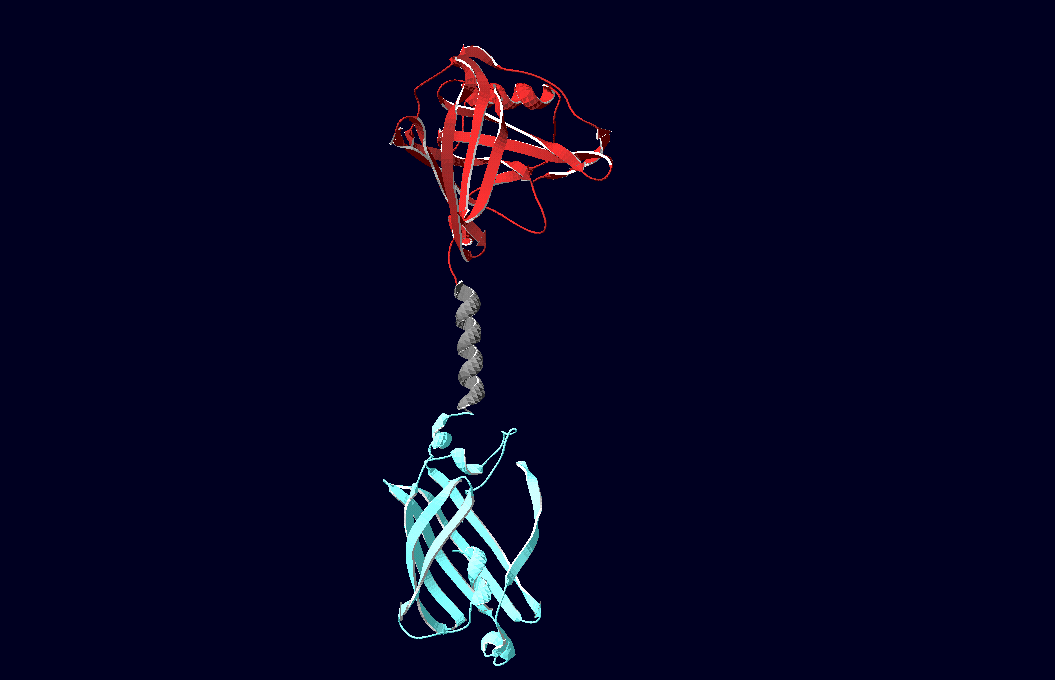
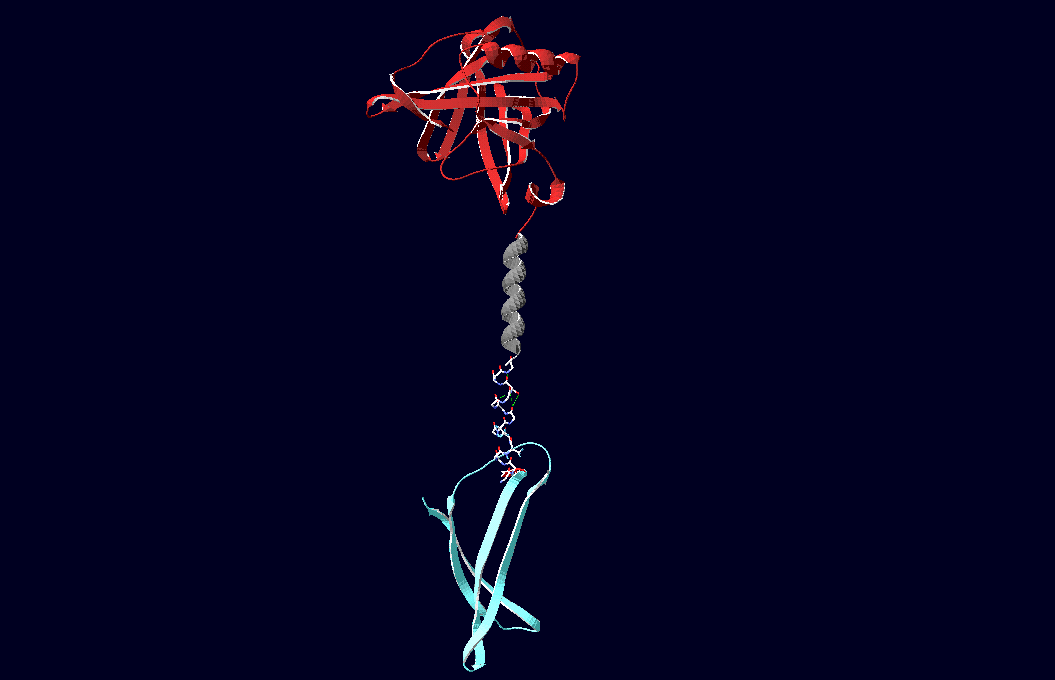
Planning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using "Nano-Engineer", Pymol and SwissPdbViewer (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.<br> | Planning the input pattern on the DNA-Origami surface, we generated various 3D-models of the huge molecule using "Nano-Engineer", Pymol and SwissPdbViewer (both freeware). Some of those models are shown here to give you an impression of the molecules we have engineered this year.<br> | ||
| - | [[Image:Freiburg2008_Fab_on_Origami_animated.gif|left|thumb|800 px|'''Fig.1:''' Some of the oligonucleotides that shape the single-stranded template fused to NIP-molecules and an anti-NIP- | + | [[Image:Freiburg2008_Fab_on_Origami_animated.gif|left|thumb|800 px|'''Fig.1:''' Some of the oligonucleotides that shape the single-stranded template fused to NIP-molecules and an anti-NIP-Fv-Fragment]] |
<br><br><br> | <br><br><br> | ||
<table> | <table> | ||
Revision as of 22:06, 29 October 2008
 "
"