Team:Freiburg/Project
From 2008.igem.org
m |
m |
||
| (57 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
<br> | <br> | ||
<br> | <br> | ||
| - | <h2>''' | + | <h2>'''Summary'''</h2> |
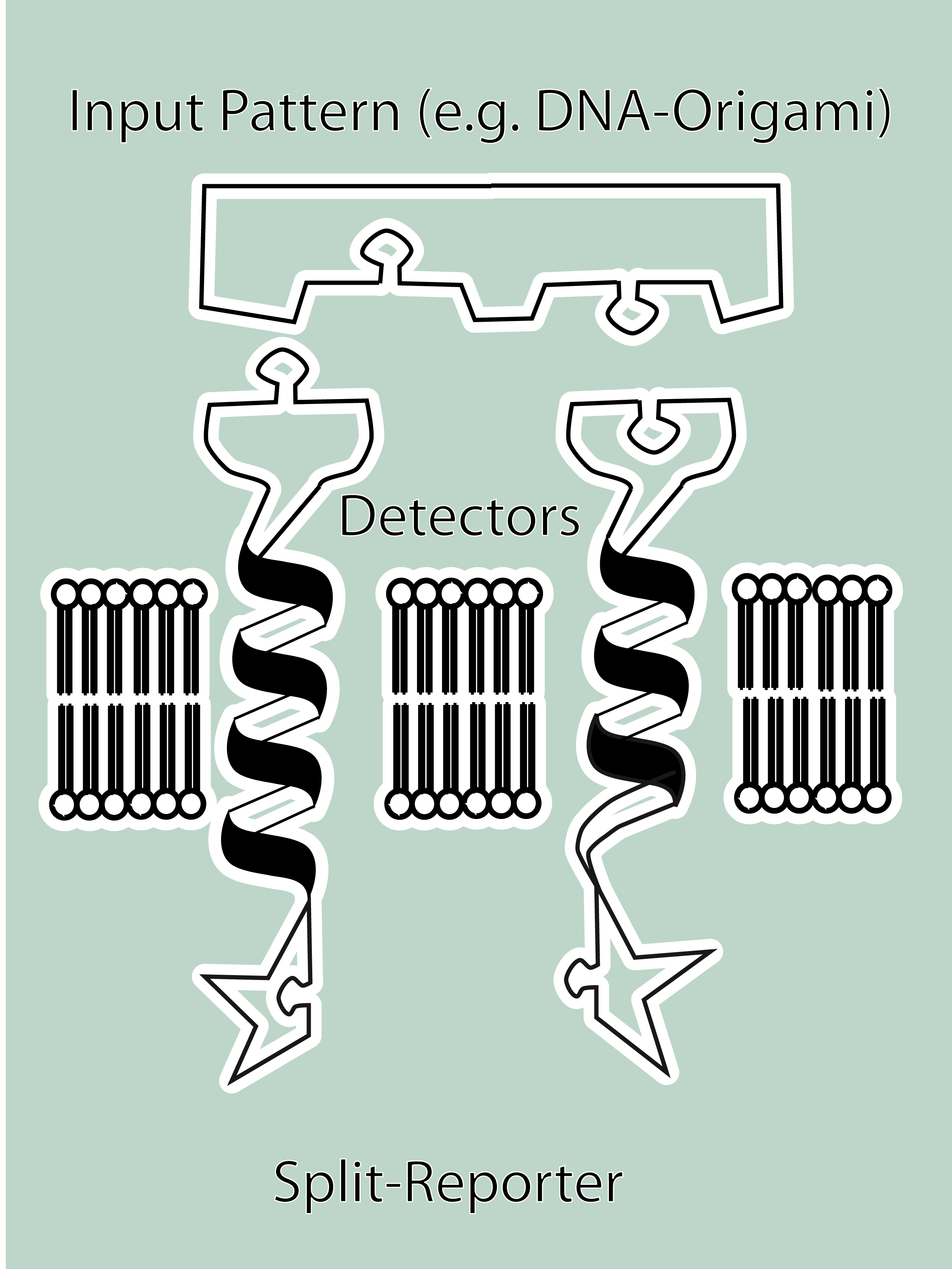
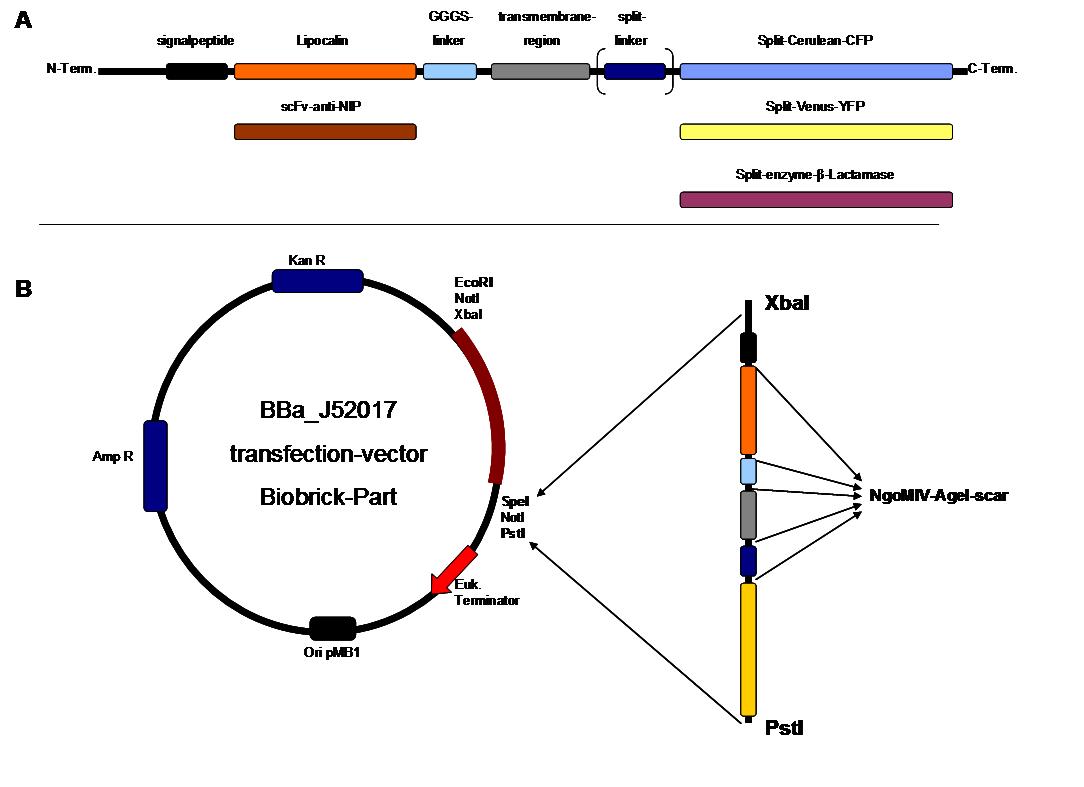
| - | + | '''Modular Synthetic Receptor System'''<br> | |
| - | + | [[Image:Freiburg08 MSRS Schema_txt.png|thumb|right|250 px]] | |
| - | + | In the following we provide a summary of our project and display the highlights of our achievements. For a more detailed view please see the reports of each subproject.<br><br> | |
| - | + | ||
| - | + | The general goal of the Freiburg 2008 team is to establish a synthetic transmembrane receptor system comprising the extracellular input devices, the extracellular receiver domain, the transducer to the cytosol and the reporter/executor in the cytosol.<br> | |
| - | the | + | As controllable receptor activation event, we chose spatial control of dimerization, which is a mechanism often used by nature. To achieve spatial control in the nanometer range we chose DNA origami for exact and complex patterns with several ligands and we opted for chemical coupling of one type of ligand to a scaffold protein for simple multimerization.<br> |
| - | + | ||
| - | <br> | + | To facilitate binding, we used a single-chain Fv antibody fragment and a designed anticalin. Both bind haptens (nitro-jodo-phenol and fluorescein) which can be conjugated to specific oligonucleotides of the DNA origami or scaffold proteins. Transduction to the cytosol is mediated by a single transmembrane helix taken either from the EGF-receptor or B-cell-receptor.<br> |
| - | + | ||
| - | + | As intracellular reporters we employ split proteins. Here, we used the split lactamase from the iGEM2007 Freiburg project as well as the split fluorescent proteins CFP/cerulean and YFP/venus. To provide steric flexibility and reach of our binding and reporting modules we interspersed linkers between each functional domain.<br> | |
| - | + | ||
| - | + | All parts submitted feature full BioBrick compatibility and in addition allow for the construction of fusion proteins due to extended Biobrick 3.0 (Freiburg) standard. Cloning was done in <i>E. coli</i>. However, we test our receptors in human or mouse cells/chassis. Thus, to direct our constructs to the cell surface we attached a eukaryotic signal peptide taken from hEGF-R to the N-terminus of each synthetic receptor. In addition to our designed synthetic receptors, we initially also employed an existing T-cell line.<br> | |
| - | + | ||
| - | + | To reach our goal within the short given time frame we started several subprojects in parallel. Our subprojects listed here are defined along these projects. Besides designing and cloning parts we spent quite some time on establishing and analyzing the biological experiments. Initially we tested interfacing of DNA origami with cells and the spatial control via DNA origami using the existing T-cell line expressing the scFv fused to a T-cell receptor. For T-cell receptor experiments we used Ca<sup>2+</sup> signaling as read out. Synthetic receptor activation was analyzed using the formation of fluorescent proteins or turnover of a fluorescent lactamase substrate. In our experiments we addressed the following questions:<br> | |
| - | + | ||
| - | + | * Can we design DNA origami to induce receptor multimerization?<br> | |
| - | + | * Can we improve DNA origami assembly yield by varying the staple oligonucleotide to scaffold DNA ratio?<br> | |
| - | + | * Can we find buffer conditions, which mediate cell viability and origami stability?<br> | |
| - | + | * Can we downsize a Ca<sup>2+</sup> release assay to fit the sample sizes of our precious DNA origami?<br> | |
| + | * Do our constructs express and then localize in the membrane of eukaryotic cells?<br> | ||
| + | * Do DNA origamis bind to cells?<br> | ||
| + | * Can we detect synthetic receptor activation?<br> | ||
| + | |||
| + | In short, we successfully addressed the first six questions. Experiments to demonstrate synthetic receptor activation are ongoing. | ||
| + | |||
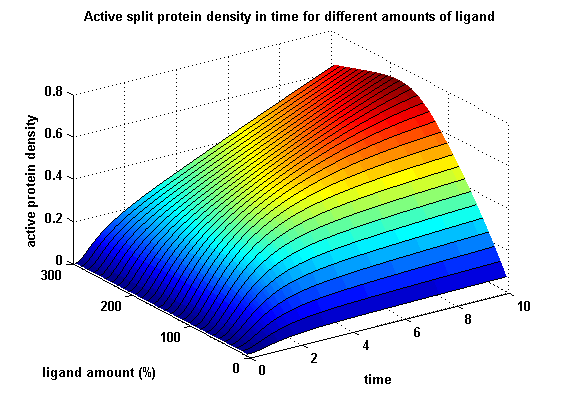
| + | For our modeling analyses we constructed various sets of differential equations describing our synthetic receptors and predicted split protein activation behaviour. | ||
| + | |||
| + | The labs of Kristian Müller and Katja Arndt provided all basic technology and support to get started. For advanced analyses several labs of the University of Freiburg and the Max-Planck-Institute of Immunobiology granted access to their instrumentation (e.g. atomic force micoscopy, confocal microscopy, FACS). | ||
<br><br> | <br><br> | ||
| + | <h2>'''Subprojects'''</h2> | ||
| + | *[[Team:Freiburg/Modeling|Modeling]]<br> | ||
| + | *[[Team:Freiburg/3D-Modeling|3D-Modeling]]<br> | ||
| + | *[[DNA-Origami|DNA-Origami]]<br> | ||
| + | *[[Team:Freiburg_Cloning Strategy|Cloning Strategy]]<br> | ||
| + | *[[Team:Freiburg_Transfection and Synthetic Receptor|Transfection and Synthetic Receptor Activation]]<br> | ||
| + | *[[Team:Freiburg_Calcium Imaging|Cell Stability, Ca<sup>2+</sup> Signaling and DNA-Origami-Binding]]<br><br> | ||
| - | + | <h2>'''Highlights'''</h2> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | <h2>'''Highlights | + | |
<table> | <table> | ||
<tr> | <tr> | ||
<td> | <td> | ||
| - | [[Team:Freiburg/Modeling|Modeling]]< | + | <h4>[[Team:Freiburg/Modeling|Modeling]]</h4> |
| - | + | <!-- Dimerization of extracellular receptor domains is an important necessity for the functionality of our Modular Synthetic Receptor System. Presenting the system a stimulus in form of spatially arranged ligands (nitro-iodo-phenol or fluorescein molecules) results in dimerization and thus the corresponding intracellular parts such as the split lactamase halves or split fluorescent proteins complement to measureable output. --> | |
| + | |||
| + | Receptor dimerization and activation was analyzed by introducing and discussing two receptor dimerization models: One T cell receptor model and one general receptor model. A proper model for the Modular Synthetic Receptor System was constructed. It consists of 10 reaction kinetic equations, 9 ordinary differential equations including 25 variable parameters. Matlab m-files are embedded for further inspection. | ||
</td> | </td> | ||
<td> | <td> | ||
| Line 50: | Line 60: | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
---- | ---- | ||
<table> | <table> | ||
<tr> | <tr> | ||
<td> | <td> | ||
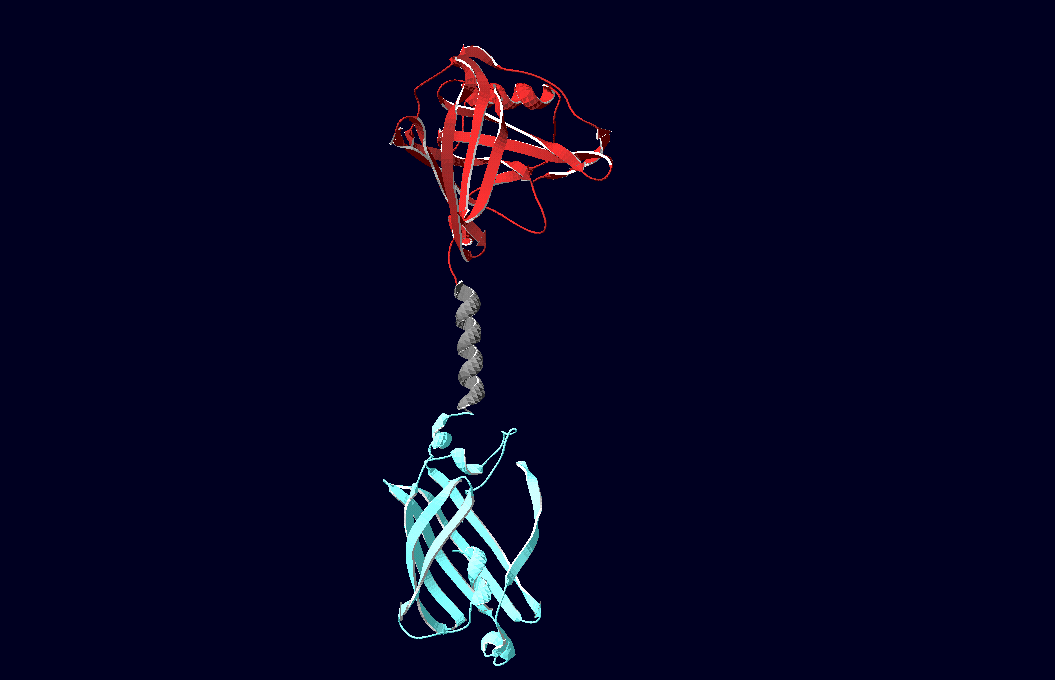
| - | [[Team | + | [[Image:Team-Freiburg2008_Lipo_alpha_nCFP.png|thumb|left|250 px|Structural model of expression part Bba_K157037]] |
| - | + | ||
| - | + | ||
</td> | </td> | ||
<td> | <td> | ||
| - | [[ | + | <h4>[[Team:Freiburg/3D-Modeling|3D-Modeling]]</h4> |
| + | We have created various three-dimensional models of our constructs using Pymol and SwissPdbViewer. Pdb-files of the huge DNA-Origami molecule were created as adequate as possible using Nano-Engineer V1.0 and then edited in Pymol. These Models were used to plan the spatial arrangement and, thus, input-pattern of antigens at a nanometer scale.<br><br><br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
| - | |||
---- | ---- | ||
<table> | <table> | ||
<tr> | <tr> | ||
<td> | <td> | ||
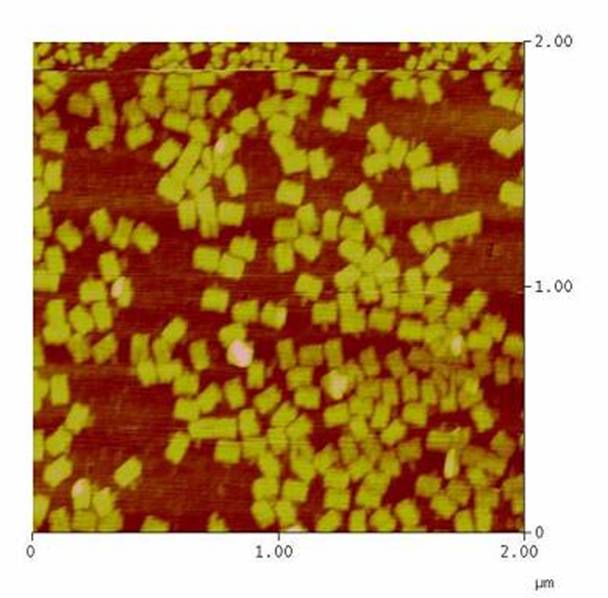
| - | [[ | + | <h4>[[DNA-Origami|DNA-Origami]]</h4> |
| + | The creation of DNA-Origami was one of the first sub-projects we engaged in. Structural planing of the input pattern on the Origami-surface and order of the modified oligo-nucleotides were the first issues, increasing of the Origami-yield and measurement of the molecule´s stability in cell culture media the later ones.<br> | ||
</td> | </td> | ||
<td> | <td> | ||
| - | [[Image: | + | [[Image:Team Freiburg2008-Origami 1zu5.jpg|thumb|right|200 px|AFM-measurement of DNA-Origami]] |
</td> | </td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| - | |||
---- | ---- | ||
<table> | <table> | ||
<tr> | <tr> | ||
<td> | <td> | ||
| - | [[ | + | [[Image:Freiburg2008 Konstrukte.jpg|thumb|left|250px|Cloning scheme]] |
| - | + | ||
</td> | </td> | ||
<td> | <td> | ||
| - | [[ | + | <h4>[[Team:Freiburg_Cloning Strategy|Cloning Strategy]]</h4> |
| + | All of our constructs were cloned successfully using our extended pre- and suffix and plasmid "pMA" (part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157000 Bba_K157000]). The broad combinatorial range given by the modular concept was almost fully exploited, so that we ended up with the submission of 13 basic and 28 composite parts.<br> | ||
| + | Once a construct was completed, it was cloned into the CMV-promoted transfection vector (part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157000 Bba_K157040]) and used for transfection of 293T-cells.<br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 94: | Line 101: | ||
<tr> | <tr> | ||
<td> | <td> | ||
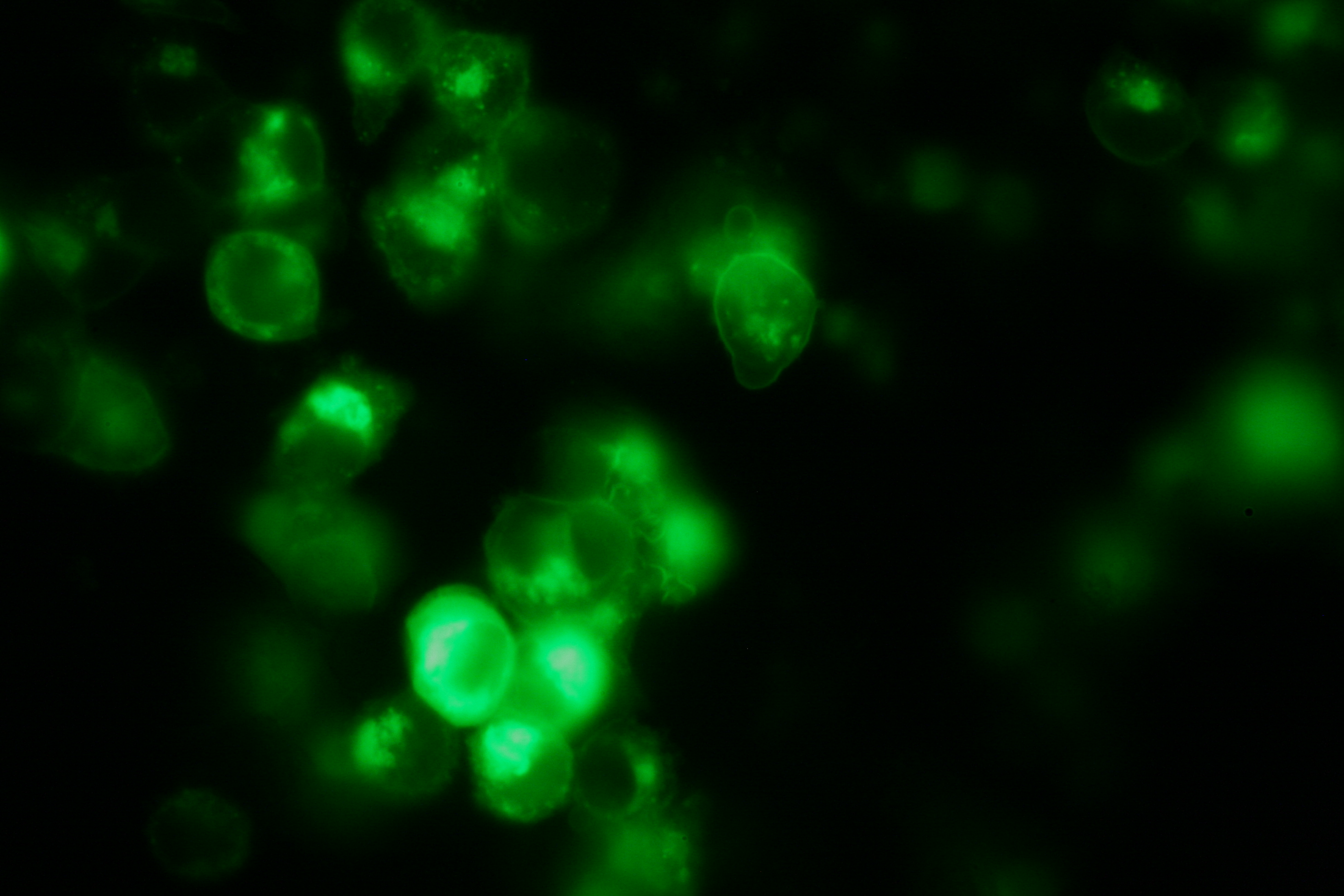
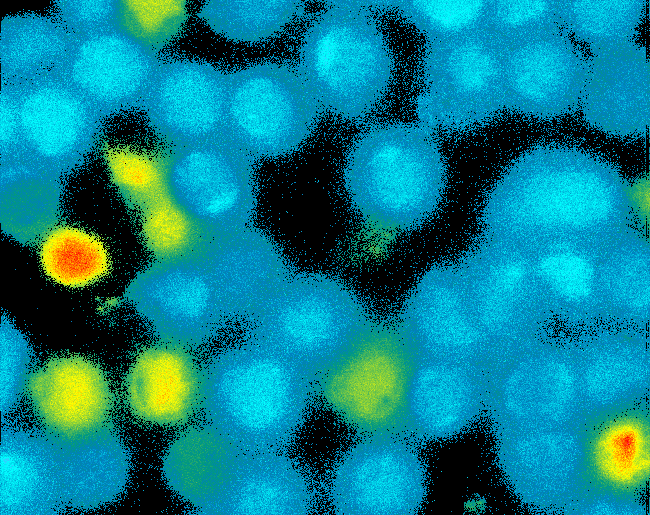
| - | [[Team: | + | <h4>[[Team:Freiburg_Transfection and Synthetic Receptor|Transfection and Synthetic Receptor Activation]]</h4> |
| + | Transfection of 293T-cells has been shown to work for most of our fusion proteins; so far, we could even show that at least our constructs with lipocalin FluA as extracellular domaine are integrated into the membrane.<br> | ||
| + | All transfections were carried out with part [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157040 Bba_K157040], one of our composite parts consisting of the transfection vector Bba_J52017 and a CMV-promotor. Due to the limited time range we were not yet able to demonstrate activation of the various possible "receptor"-pairs, but we are hoping to receive a positive result soon.<br> | ||
</td> | </td> | ||
<td> | <td> | ||
| - | [[Image: | + | [[Image:Freiburg2008_Fluomembrane.jpg|thumb|right|200px|Membrane-localization of Part Bba_K157032-YFP]] |
</td> | </td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
| + | ---- | ||
| + | <table> | ||
| + | <tr> | ||
| + | <td> | ||
| + | [[Image:TeamFreiburg2008_Pervanadate1.png|thumb|left|200px|Ca<sup>2+</sup>-influx measurement]] | ||
| + | </td> | ||
| + | <td> | ||
| + | <h4>[[Team:Freiburg_Calcium Imaging|Cell Stability, Ca<sup>2+</sup> Signaling and DNA-Origami-Binding]]</h4> | ||
| + | Due to the small total amounts of DNA-Origami we could not use FACS to measure calcium influx and, thus, T-cell activation. The alternative we found were "Nano-Slides" for inverse fluorescence microscopy; T-cells were immobilized on the poly-L-Lysine coated slides and stimulated under the microscope allowing for real-time observation of calcium-influx. Binding of fluorescent labeled DNA origami to cells was analyzed by confocal microscopy.<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| - | <h2>''' | + | <h2>'''[[Image:MO2.jpg|50px|]]Literature'''</h2> |
'''Split-fluorophores:'''<br> | '''Split-fluorophores:'''<br> | ||
| - | + | *Chang-Deng Hu, Yurii Chinenov, Tom K. Kerppola: ”Visualization of Interactions among bZIP and Rel Family Proteins in Living Cells Using Bimolecular Fluorescence Complementation”, Molecular Cell, Vol. 9, 789–798, April, 2002<br> | |
| - | + | *Chang Deng Hu, Tom K. Kerppola: “Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis”, Nat Biotechnol. 2003 May; 21(5):539-545 (doi:10. 1038/nbt816)<br> | |
| - | + | *Tom K. Kerppola: “Design and implementation of bimolecular fluorescence complementation (BiFC) assays for the visualization of protein interactions in living cells”, Nat Protoc. 2006;1(3):1278-1286 (doi:10.1038/nprot.2006.201)<br> | |
| - | + | *Nagai, T. et al. “A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications” J. Biol. Chem. 276, 29188-29194, 2001<br> | |
| - | + | *Roger Y. Tsien et al. „Creating new fluorescent probes for cell biology“, Nature Biotechnology Reviews, Vol. 3, 906-918, 2002<br> | |
'''LipocalinFluA:'''<br> | '''LipocalinFluA:'''<br> | ||
| - | + | *Gerald Beste, Frank S. Schmidt, Thomas Stibora and A. Skerra: “Small antibody-like proteins with prescribed ligand specificities derived from the lipocalin fold“,Proc. Natl. Acad. Sci. USA Vol. 96, pp. 1898–1903, March 1999 Biochemistry<br> | |
| - | + | *Ingo P. Korndörfer, Gerald Beste and A. Skerra: “Crystallographic Analysis of an “Anticalin” With Tailored Specificity for Fluorescein Reveals High Structural Plasticity of the Lipocalin Loop Region”, PROTEINS: Structure, Function, and Bioinformatics 53:121–129 (2003)<br> | |
'''DNA-Origami:'''<br> | '''DNA-Origami:'''<br> | ||
| - | Paul W. K. Rothemund: Nature 440, 297-302 (16 March 2006)<br> | + | *Paul W. K. Rothemund: "Folding DNA to create nanoscale shapes and patterns", Nature 440, 297-302 (16 March 2006)<br> |
'''Antibody B1-8:'''<br> | '''Antibody B1-8:'''<br> | ||
| - | + | *Ana Cumano and Klaus Rajewski: “Clonal recruitment and somatic mutation in the generation of immunological memory to the hapten NP”, The EMBO Journal vol. 5 no.10 pp. 2459-2468, 1986<br> | |
| - | + | *D. Allen, T. Simon, F. Sablitzky, K. Rajewski and A. Cumano: “Antibody engineering for the analysis of affinity maturation of an anti-hapten response”, The EMBO Journal vol. 7 no.7 pp. 1995-2001, 1988<br> | |
'''Modeling:'''<br> | '''Modeling:'''<br> | ||
| - | + | *Martin F. Bachmann, Michael Salzmann, Annette Oxenius and Pamela S. Ohashi: "Formation of TCR dimers/trimers as a crucial step for T cell activation", Eur. J. Immunol., 1998<br> | |
| - | + | *Martin F. Bachmann and Pamela S. Ohashi: "The role of T-cell receptor dimerization in T-cell activation", Review Immunology Today, 1999<br> | |
| - | + | *João Sousa and Jorge Carneiro: "A mathematical analysis of TCR serial triggering and down-regulation", Eur. J. Immunol., 2000<br> | |
'''TCR activation:'''<br> | '''TCR activation:'''<br> | ||
| - | + | *Susana Minguet, Mahima Swamy, Balbino Alarcón, Immanuel F. Luescher and Wolfgang W.A. Schamel: "Full Activation of the T Cell Receptor requires Both Clustering and Conformational Changes at CD3", Immunity, 2006 | |
}} | }} | ||
Latest revision as of 08:24, 30 October 2008
|
Project Report |
_project report
SummaryModular Synthetic Receptor System In the following we provide a summary of our project and display the highlights of our achievements. For a more detailed view please see the reports of each subproject. The general goal of the Freiburg 2008 team is to establish a synthetic transmembrane receptor system comprising the extracellular input devices, the extracellular receiver domain, the transducer to the cytosol and the reporter/executor in the cytosol. To facilitate binding, we used a single-chain Fv antibody fragment and a designed anticalin. Both bind haptens (nitro-jodo-phenol and fluorescein) which can be conjugated to specific oligonucleotides of the DNA origami or scaffold proteins. Transduction to the cytosol is mediated by a single transmembrane helix taken either from the EGF-receptor or B-cell-receptor. As intracellular reporters we employ split proteins. Here, we used the split lactamase from the iGEM2007 Freiburg project as well as the split fluorescent proteins CFP/cerulean and YFP/venus. To provide steric flexibility and reach of our binding and reporting modules we interspersed linkers between each functional domain. All parts submitted feature full BioBrick compatibility and in addition allow for the construction of fusion proteins due to extended Biobrick 3.0 (Freiburg) standard. Cloning was done in E. coli. However, we test our receptors in human or mouse cells/chassis. Thus, to direct our constructs to the cell surface we attached a eukaryotic signal peptide taken from hEGF-R to the N-terminus of each synthetic receptor. In addition to our designed synthetic receptors, we initially also employed an existing T-cell line. To reach our goal within the short given time frame we started several subprojects in parallel. Our subprojects listed here are defined along these projects. Besides designing and cloning parts we spent quite some time on establishing and analyzing the biological experiments. Initially we tested interfacing of DNA origami with cells and the spatial control via DNA origami using the existing T-cell line expressing the scFv fused to a T-cell receptor. For T-cell receptor experiments we used Ca2+ signaling as read out. Synthetic receptor activation was analyzed using the formation of fluorescent proteins or turnover of a fluorescent lactamase substrate. In our experiments we addressed the following questions:
In short, we successfully addressed the first six questions. Experiments to demonstrate synthetic receptor activation are ongoing. For our modeling analyses we constructed various sets of differential equations describing our synthetic receptors and predicted split protein activation behaviour. The labs of Kristian Müller and Katja Arndt provided all basic technology and support to get started. For advanced analyses several labs of the University of Freiburg and the Max-Planck-Institute of Immunobiology granted access to their instrumentation (e.g. atomic force micoscopy, confocal microscopy, FACS).
Subprojects
Highlights
|
 "
"