Team:Freiburg Cloning Strategy
From 2008.igem.org
| (14 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
<br> | <br> | ||
| - | + | <h2>Introduction</h2> | |
| - | To form different types of synthetic receptor constructs a | + | To form different types of synthetic receptor constructs a modular building set was used. We designed the genes of two parts coding for extracellular binding proteins (anti NIP scFv, anti-Flu Lipocalin) and four different components of an intracellular signal transduction reporter protein (Cerulean CFP, Venus YFP, β-Lactamase, and Luciferase). These reporters were designed as split-proteins to achieve activaton only when two receptors come together and form a cluster. With this strategy, a signal is only given by two binding-molecules that are next to each other as it is realized in the DNA-origami structure or the NIP and Fluorescein-coupled BSA. <br> |
| - | The whole receptor construct consists out of a | + | The whole receptor construct consists out of (i) a signal peptide, (ii) an extracellular receptor domain, (iii) a GGGS-linker followed by (iv) a transmembrane region and (v) an intracellular split-reporter-protein. The signal peptide sequence equal to that of human EGF-receptor erbb1 ensures the transport of the protein construct to the cytoplasma membrane. A GGGS-linker is necessary to create a distance between membrane and the recognition-site to tower cell surface structures as glycoproteins and glycolipids. The sequence of the transmembrane region is taken from the EGF-receptor erbb1 as well. The C-terminal split-parts of Venus-YFP and Cerulean CFP were not directly fused to the transmembrane region. To get a little bit more flexibility in binding to the N-terminal part, a split fluorophor-linker was assembled in between.<br><br> |
| - | Almost all parts were ordered by | + | |
| - | The | + | Almost all parts were ordered by gene synthesis in a pMA-vector system that is adapted for cloning BioBrick parts. The constructs were designed according to BioBrick 3.0 standard with the modification for fusion-proteins proposed by the iGEM Freiburg Team 2007 ([https://2007.igem.org/Freiburg07/report_fusion_parts FreiGEM07_report_fusion_part]). Keeping the standard iGEM prefix and suffix, the restriction sites NgoMIV and AgeI were added to create functional fusions without stop-codons. <br> |
| + | All possible variations of the receptor constructs were fused together in the pMA-vector and afterwards inserted into a transfection vector. The transfection vector is a BioBrick 2006 part from the Ljubljana, Slovenia team ([http://partsregistry.org/Part:BBa_J52017 BBa_J52017]) and includes a kanamycin and ampicillin resistance cassette. There is a pMB1 Ori and the multiple cloning site consist of the BioBrick Prefix and Suffix restriction enzymes (Prefix: EcoRI, NotI, XbaI/ Suffix: SpeI, NotI, PstI) followed by an eukaryotic terminator sequence. To achieve a strong expression, a constitutive promoter construct was needed. Because of this, the Cytomegalovirus [CMV] promotor was amplified by PCR using the BioBrick part [http://partsregistry.org/Part:BBa_J52038 BBa_J52038 ]from the 2006 Ljubljana, Slovenia team as well and primers mentioned below. The forward primer bound in the iGEM-prefix, while the reverse primer bound at the end of the CMV-coding region. The reverse primer also contained an overhang to get the iGEM-suffix directly to the end of the promoter sequence which later allowed the cloning into the transfection vector. The CMV promoter was cloned into the vector by using the EcoRI and PstI restriction sites. To test the expression activity of the vector with the CMV promoter, the gene of the yellow fluorescent protein [YFP] was cloned downstream of the promoter region by using XbaI, PstI restriction enzymes for the YFP insert and SpeI, PstI for the vector to open the Biobrick suffix. The transfection of the resulting plasmid into 293T cells shows full functionality.<br> | ||
An overview about the cloning is given in table 1<br> | An overview about the cloning is given in table 1<br> | ||
<br><br> | <br><br> | ||
| Line 18: | Line 19: | ||
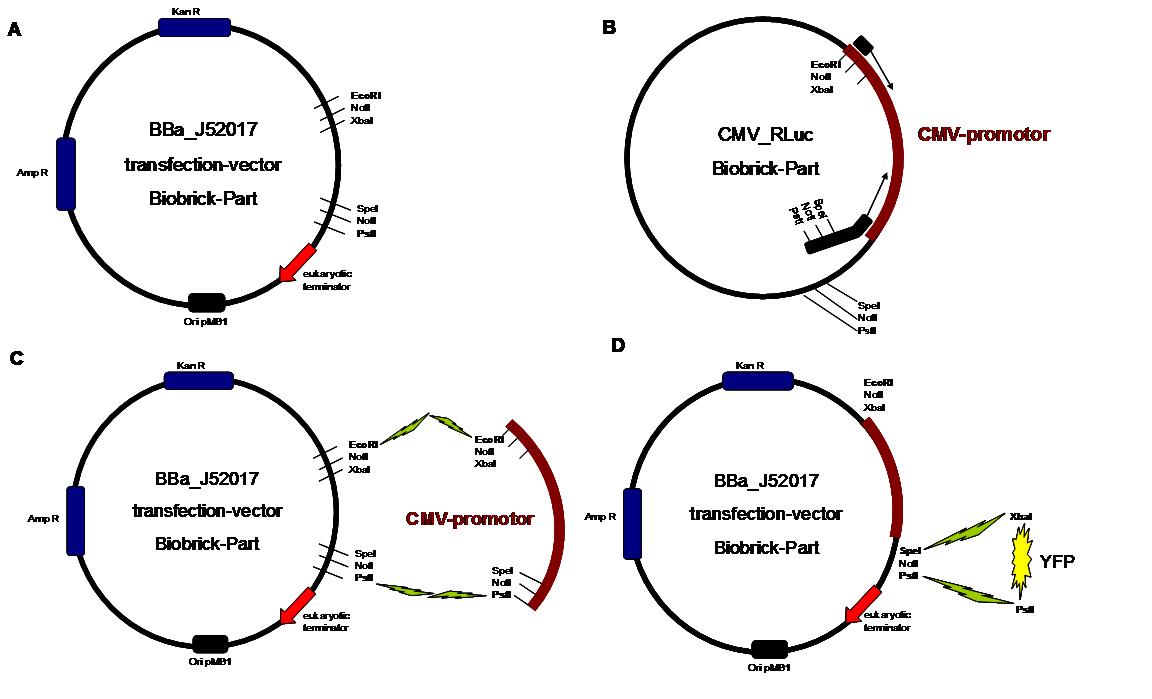
<span style="font-weight: bold;">Figure 1_cloning strategy</span> | <span style="font-weight: bold;">Figure 1_cloning strategy</span> | ||
. <span style="font-weight: bold;">A</span> | . <span style="font-weight: bold;">A</span> | ||
| - | <small>shows the Biobrick part BBa_J52017. The | + | <small>shows the Biobrick part BBa_J52017. The transfection vector |
| - | for eukaryotic cell systems has an ampicillin | + | for eukaryotic cell systems has an ampicillin and a kanamycin |
resistance cassette. The multiple cloning site contains the Biobrick | resistance cassette. The multiple cloning site contains the Biobrick | ||
standard restriction sites EcoRI, NotI, XbaI, SpeI, NotI, PstI followed | standard restriction sites EcoRI, NotI, XbaI, SpeI, NotI, PstI followed | ||
| - | by | + | by a eukaryotic terminator sequence. <big><span |
style="font-weight: bold;">B</span></big> The | style="font-weight: bold;">B</span></big> The | ||
| - | CMV | + | CMV promotor fragment was obtained by PCR from the Biobrick BBa-J52038 |
template. <big><span style="font-weight: bold;">C</span></big> | template. <big><span style="font-weight: bold;">C</span></big> | ||
| - | The PCR product was cloned into the | + | The PCR product was cloned into the transfection vector by EcoRI and |
| - | PstI to get a final eukaryotic transfection | + | PstI to get a final eukaryotic transfection system. <big><span |
style="font-weight: bold;">D</span></big> To | style="font-weight: bold;">D</span></big> To | ||
| - | test the efficiency of expression a gene | + | test the efficiency of expression, a gene fragment coding for the yellow |
| - | fluorescent protein was | + | fluorescent protein was cloned into the vector downstream of the CMV Promotor</small>. |
<br><br> | <br><br> | ||
| Line 38: | Line 39: | ||
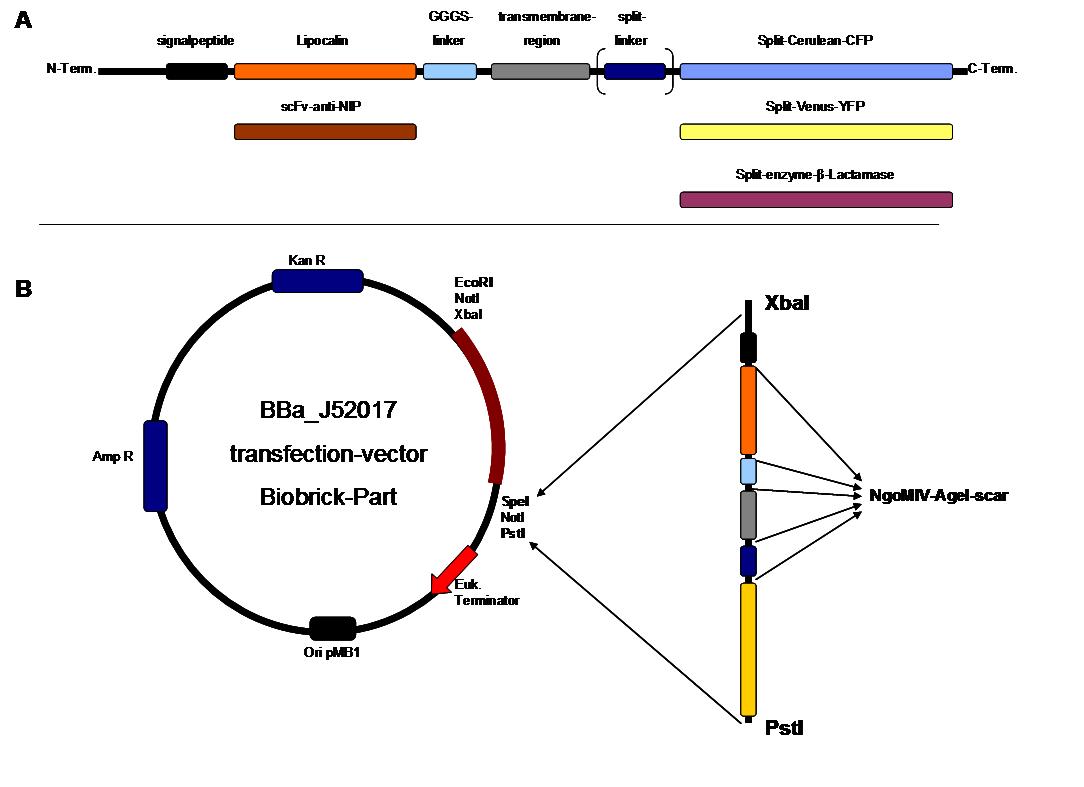
. <span style="font-weight: bold;">A</span> | . <span style="font-weight: bold;">A</span> | ||
<small>figure 2 A gives an overview about the cloning constructs. | <small>figure 2 A gives an overview about the cloning constructs. | ||
| - | The N-terminal signal | + | The N-terminal signal peptide ensures protein transport to the |
| - | + | cytoplasma membrane. Lipocalin and the scFv-anti-NIP are the | |
| - | extracytoplasmatic parts of the construct to mediate | + | extracytoplasmatic parts of the construct to mediate signal transduction |
into the cell. The GGGS-Linker keeps a distance to the | into the cell. The GGGS-Linker keeps a distance to the | ||
| - | + | transmembrane region to overcome surface structures of the cell and to | |
| - | avoid a total inflexibility. | + | avoid a total inflexibility. The split fluorophor linker is only necessary |
| - | for the C-terminal | + | for the C-terminal split parts of Cerulean-CFP and |
| - | + | Venus-YFP. The split enzymes β-Lactamase and Luciferase | |
and the split-fluorophors CFP and YFP are the cytoplasmatic parts of | and the split-fluorophors CFP and YFP are the cytoplasmatic parts of | ||
the constructs. If there is a clustering of this synthetic | the constructs. If there is a clustering of this synthetic | ||
receptor-system caused by the corresponding binding parts of Lipocalin | receptor-system caused by the corresponding binding parts of Lipocalin | ||
| - | and scFv-anti-NIP the split parts come together to create a functional | + | and scFv-anti-NIP, the split parts come together to create a functional |
| - | protein, which allows | + | protein, which allows detection. </small><br> |
<span style="font-weight: bold;">B</span> <small>The | <span style="font-weight: bold;">B</span> <small>The | ||
different constructions described in figure 2.A were cloned into the | different constructions described in figure 2.A were cloned into the | ||
| - | + | transfection vector system by using the restriction sites XbaI and PstI | |
to ensure a functional ATG-start codon which is part of the XbaI | to ensure a functional ATG-start codon which is part of the XbaI | ||
recognition-sequence in the iGEM-prefix.</small> | recognition-sequence in the iGEM-prefix.</small> | ||
| Line 63: | Line 64: | ||
style="" lang="EN-GB">overview about<b style=""> | style="" lang="EN-GB">overview about<b style=""> | ||
</b>the cloning steps to create the different types of synthetic | </b>the cloning steps to create the different types of synthetic | ||
| - | receptors. To get further information about the | + | receptors. To get further information about the composite parts see </span></small></p> |
<p class="MsoNormal"><span style="" lang="EN-GB"><small>https://2008.igem.org/Team:Freiburg/Parts</small> | <p class="MsoNormal"><span style="" lang="EN-GB"><small>https://2008.igem.org/Team:Freiburg/Parts</small> | ||
<br><br> | <br><br> | ||
| Line 987: | Line 988: | ||
<br> | <br> | ||
| - | + | <h2>Methods</h2> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
The cloning was started with a preparative digestion of the DNA-Plasmids. To clone fusion parts the vector constructs were digested with AgeI and PstI to open the Biobrick suffix. The inserts were digested with NgoMIV and PstI. For cloning into the transfection-vector the enzymes SpeI and PstI were used for vector and XbaI, PstI for insert to keep up the ATG-start codon in the XbaI restriction site of the biobrick suffix. All restriction-enzymes were ordered from New England Biolabs. After digestion the DNA-fragments were separated on a 1% agarose gel. The DNA-band of interest was isolated and purified with the QIAGEN QIAquick Gel Extraction Kit. For the ligation a 3 molar excess of the insert was put together with the vector-fragment and ligated with a Quick ligase (New England Biolabs). After half an our at room temperature the DNA was transformed to chemical competent E.coli strain XL1 cells, plated on 2YT-agar-plates and incubated at 37°C over night. After picking clones and growing in 5ml LB-medium, the plasmid DNA was isolated by QIAGEN QIAprep Spin Miniprep Kit. A test digestion was prepared with about 0,5µg Plasmid DNA and NotI restriction enzyme to isolate the fusion-protein from the vector and to control if the expected bands were obtained. After a positive result the clones were sent to GATC-Biotech for sequencing. | The cloning was started with a preparative digestion of the DNA-Plasmids. To clone fusion parts the vector constructs were digested with AgeI and PstI to open the Biobrick suffix. The inserts were digested with NgoMIV and PstI. For cloning into the transfection-vector the enzymes SpeI and PstI were used for vector and XbaI, PstI for insert to keep up the ATG-start codon in the XbaI restriction site of the biobrick suffix. All restriction-enzymes were ordered from New England Biolabs. After digestion the DNA-fragments were separated on a 1% agarose gel. The DNA-band of interest was isolated and purified with the QIAGEN QIAquick Gel Extraction Kit. For the ligation a 3 molar excess of the insert was put together with the vector-fragment and ligated with a Quick ligase (New England Biolabs). After half an our at room temperature the DNA was transformed to chemical competent E.coli strain XL1 cells, plated on 2YT-agar-plates and incubated at 37°C over night. After picking clones and growing in 5ml LB-medium, the plasmid DNA was isolated by QIAGEN QIAprep Spin Miniprep Kit. A test digestion was prepared with about 0,5µg Plasmid DNA and NotI restriction enzyme to isolate the fusion-protein from the vector and to control if the expected bands were obtained. After a positive result the clones were sent to GATC-Biotech for sequencing. | ||
| - | The GGGS-Linker was produced by Klenow -fill-in-PCR. Two primers were designed align to each other at 60°C and filled to a complete | + | The GGGS-Linker was produced by Klenow -fill-in-PCR. Two primers were designed align to each other at 60°C and filled to a complete double-strand by Klenow Polymerase fragment.<br> |
<br> | <br> | ||
| - | Digestion Protocol < | + | <h4>Digestion Protocol</h4> |
- about 2µg Plasmid-Prep in 20µl <br> | - about 2µg Plasmid-Prep in 20µl <br> | ||
- 2 µl NEB Buffer 10x<br> | - 2 µl NEB Buffer 10x<br> | ||
| Line 1,006: | Line 1,001: | ||
- 0.2µl BSA 100x<br> | - 0.2µl BSA 100x<br> | ||
<br> | <br> | ||
| - | Ligation< | + | <h4>Ligation</h4> |
- 10µl volume of vector and insert DNA (about 50ng vector-DNA)<br> | - 10µl volume of vector and insert DNA (about 50ng vector-DNA)<br> | ||
- 1 µl DNA Quick Ligase (New England Biolabs)<br> | - 1 µl DNA Quick Ligase (New England Biolabs)<br> | ||
- 10 µl Quick Ligase Buffer<br> | - 10 µl Quick Ligase Buffer<br> | ||
<br> | <br> | ||
| - | Analytic digestion< | + | <h4>Analytic digestion</h4> |
- about 0.5 µg Plasmid-DNA in 5µl<br> | - about 0.5 µg Plasmid-DNA in 5µl<br> | ||
- 5µl H2O<br> | - 5µl H2O<br> | ||
| Line 1,018: | Line 1,013: | ||
- 0.1 µl BSA<br> | - 0.1 µl BSA<br> | ||
<br> | <br> | ||
| - | Transformation< | + | <h4>Transformation</h4> |
- Competent cells (100µl) werde defrosted on ice<br> | - Competent cells (100µl) werde defrosted on ice<br> | ||
- 10µl of the ligation was added<br> | - 10µl of the ligation was added<br> | ||
| Line 1,029: | Line 1,024: | ||
- cells werde plated on 2YT-agar-plates with antibiotics<br> | - cells werde plated on 2YT-agar-plates with antibiotics<br> | ||
<br> | <br> | ||
| - | Klenow fill in reaction< | + | <h4>Klenow fill in reaction</h4> |
- 25pmol forward primer<br> | - 25pmol forward primer<br> | ||
- 25pmol reverse primer<br> | - 25pmol reverse primer<br> | ||
| Line 1,039: | Line 1,034: | ||
| - | + | <h2>Results</h2> | |
| + | |||
| + | All composite parts were succesfully cloned and controlled by sequencing and transfection as well ([https://2008.igem.org/Team:Freiburg_Transfection_and_Synthetic_Receptor Freiburg Transfection and Synthetic Receptor]) | ||
| + | |||
| + | <h2>Biobrick fusion protein</h2> | ||
| + | |||
| + | The present BioBrick prefix and suffix rules are not compatible with modular protein design. Thus as in 2007, we proposed an extension of the present standard for fusion proteins in which two restriction sites are added in frame adjacent to the coding sequence. The basic parts and as well all composite parts follow this strategy. | ||
| + | To get further information see [https://2007.igem.org/Freiburg07/report_fusion_parts FreiGEM07_report_fusion_part]<br> | ||
| + | |||
| + | |||
| + | <h2>Cloned parts</h2> | ||
The cloning parts scFv-anti-NIP, lipocalin-FluA, Split-Venus-N-YFP, Split-Venus-C-YFP, Cerulean-N-CFP and Cerulean-C-CFP were obtained by genesynthesis and ordered by the GENEART company. The β-Lactamase fragments were adopted from genesynthesis GENEART order in 2007. The erbb1-transmembrane, Split-Fluorophor-Linker and the erbb1-signalpeptide were synthesized by ATG:biosynthetics. The CMV promoter cloning part was produced by PCR with the template J52038 using the CMV_RLuc_Forward-primer and CMV_RLuc_Reverse-primer. To synthesize the GGGS-linker a Klenow-fill-in-reaction was performed by using Klenow-fragment without exonuclease activity and GGGS-forward and reverse primer.<br> | The cloning parts scFv-anti-NIP, lipocalin-FluA, Split-Venus-N-YFP, Split-Venus-C-YFP, Cerulean-N-CFP and Cerulean-C-CFP were obtained by genesynthesis and ordered by the GENEART company. The β-Lactamase fragments were adopted from genesynthesis GENEART order in 2007. The erbb1-transmembrane, Split-Fluorophor-Linker and the erbb1-signalpeptide were synthesized by ATG:biosynthetics. The CMV promoter cloning part was produced by PCR with the template J52038 using the CMV_RLuc_Forward-primer and CMV_RLuc_Reverse-primer. To synthesize the GGGS-linker a Klenow-fill-in-reaction was performed by using Klenow-fragment without exonuclease activity and GGGS-forward and reverse primer.<br> | ||
Latest revision as of 09:01, 16 November 2008
|
_cloning strategy
IntroductionTo form different types of synthetic receptor constructs a modular building set was used. We designed the genes of two parts coding for extracellular binding proteins (anti NIP scFv, anti-Flu Lipocalin) and four different components of an intracellular signal transduction reporter protein (Cerulean CFP, Venus YFP, β-Lactamase, and Luciferase). These reporters were designed as split-proteins to achieve activaton only when two receptors come together and form a cluster. With this strategy, a signal is only given by two binding-molecules that are next to each other as it is realized in the DNA-origami structure or the NIP and Fluorescein-coupled BSA. Almost all parts were ordered by gene synthesis in a pMA-vector system that is adapted for cloning BioBrick parts. The constructs were designed according to BioBrick 3.0 standard with the modification for fusion-proteins proposed by the iGEM Freiburg Team 2007 (FreiGEM07_report_fusion_part). Keeping the standard iGEM prefix and suffix, the restriction sites NgoMIV and AgeI were added to create functional fusions without stop-codons.
Table1_cloning strategy overview about the cloning steps to create the different types of synthetic receptors. To get further information about the composite parts see https://2008.igem.org/Team:Freiburg/Parts
Step 1 Vector digestion: EcoRI + PstI Insert
digestion: EcoRI + PstI BBa-J52017 _CMV-promotor Step
2 Vector digestion: AgeI+SpeI Insert
digestion: NgoMIV+SpeI pMA-BBFR
_ SPLIT-Linker C-YFP C-CFP Step
3 Vector digestion: AgeI+SpeI Insert
digestion: NgoMIV+SpeI pMA-BBFR
_egfR-Tm _ N-β-Lactamase _ C-β-Lactamase _ SPLIT-Linker_ C-YFP _ N-YFP _ SPLIT-Linker_ C-CFP _ N-CFP _ BB058 (Luciferase) _ BB057 (Luciferase) Step
4 Vector digestion: AgeI+SpeI Insert
digestion: NgoMIV+SpeI pMA-BBFR
_SP _scFv-anti-NIP _ Lipocalin Step
5 Vector digestion: AgeI+SpeI Insert
digestion: NgoMIV+SpeI pMA-BBFR
_SP_ scFv-anti-NIP and pMA-BBFR-+SP_ Lipocalin _GGGS-linker (produced by Klenow fill in) Step
6 Vector digestion: AgeI+SpeI Insert
digestion: NgoMIV+SpeI pMA-BBFR
_SP_ scFv-anti-NIP _ GGGS-Li and pMA-BBFR
_ SP_ Lipocalin _ GGGS-Li _
egfR-Tm _ N-β-Lactamase _
egfR-Tm _ C-β-Lactamase _
egfR-Tm _ SPLIT-Linker_ C-YFP _
egfR-Tm _ N-YFP _
egfR-Tm _ SPLIT-Linker_ C-CFP _
egfR-Tm _ N-CFP _
egfR-Tm _ BB058 (Luciferase) _
egfR-Tm _ BB057 (Luciferase) Step
7 Vector digestion:
SpeI + PstI Insert
digestion: XbaI + PstI BBa-J52017_CMV _SP_ scFv-anti-NIP_GGGS-Li_egfR-Tm_N-β-Lactamase _ SP_ scFv-anti-NI _GGGS-Li_ egfR-Tm_C-β-Lactamase _ SP_ scFv-anti-NIP_GGGS-Li_
egfR-Tm_SPLIT-Linker_C-YFP _ SP_ scFv-anti-NIP_GGGS-Li_ egfR-Tm_N-YFP _ SP_ scFv-anti-NIP_GGGS-Li_
egfR-Tm_SPLIT-Linker_C-CFP _ SP_ scFv-anti-NIP_GGGS-Li_ egfR-Tm_N-CFP _ SP_ scFv-anti-NIP_GGGS-Li _ egfR-Tm_BB058 (Luciferase) _ SP_ scFv-anti-NIP_GGGS-Li _ egfR-Tm_BB057 (Luciferase) _ SP_ Lipocalin _GGGS-Li_ egfR-Tm_N-β-Lactamase _ SP_ Lipocalin _GGGS-Li_ egfR-Tm_C-β-Lactamase _ SP_ Lipocalin _GGGS-Li_
egfR-Tm_SPLIT-Linker_ C-YFP _ SP_ Lipocalin _GGGS-Li_
egfR-Tm_N-YFP _ SP_ Lipocalin _GGGS-Li_
egfR-Tm_SPLIT-Linker_ C-CFP _ SP_ Lipocalin _GGGS-Li_
egfR-Tm_N-CFP _ SP_ Lipocalin _GGGS-Li__ egfR-Tm _ BB058 (Luciferase) _ SP_ Lipocalin _GGGS-Li__ egfR-Tm _ BB057 (Luciferase) - about 2µg Plasmid-Prep in 20µl - 10µl volume of vector and insert DNA (about 50ng vector-DNA) - about 0.5 µg Plasmid-DNA in 5µl - Competent cells (100µl) werde defrosted on ice - 25pmol forward primer All composite parts were succesfully cloned and controlled by sequencing and transfection as well (Freiburg Transfection and Synthetic Receptor)
The present BioBrick prefix and suffix rules are not compatible with modular protein design. Thus as in 2007, we proposed an extension of the present standard for fusion proteins in which two restriction sites are added in frame adjacent to the coding sequence. The basic parts and as well all composite parts follow this strategy.
To get further information see FreiGEM07_report_fusion_part The cloning parts scFv-anti-NIP, lipocalin-FluA, Split-Venus-N-YFP, Split-Venus-C-YFP, Cerulean-N-CFP and Cerulean-C-CFP were obtained by genesynthesis and ordered by the GENEART company. The β-Lactamase fragments were adopted from genesynthesis GENEART order in 2007. The erbb1-transmembrane, Split-Fluorophor-Linker and the erbb1-signalpeptide were synthesized by ATG:biosynthetics. The CMV promoter cloning part was produced by PCR with the template J52038 using the CMV_RLuc_Forward-primer and CMV_RLuc_Reverse-primer. To synthesize the GGGS-linker a Klenow-fill-in-reaction was performed by using Klenow-fragment without exonuclease activity and GGGS-forward and reverse primer. |
 "
"