Team:Bologna/Project
From 2008.igem.org
(→Ecoli.PROM: an Erasable and Programmable Genetic Memory in E. coli) |
(→Ecoli.PROM: an Erasable and Programmable Genetic Memory in E. coli) |
||
| Line 53: | Line 53: | ||
[[Image:bba020.jpg|center|thumbnail|400 px|Figure 1. Upper panel: Closed-loop configuration (K079020); lower panel Open-loop configuration (K079026)]] | [[Image:bba020.jpg|center|thumbnail|400 px|Figure 1. Upper panel: Closed-loop configuration (K079020); lower panel Open-loop configuration (K079026)]] | ||
| - | + | ||
Fluorescence images demonstrate the repression due to the presence of the Lac operator. | Fluorescence images demonstrate the repression due to the presence of the Lac operator. | ||
| - | |||
{|align="center" | {|align="center" | ||
| Line 62: | Line 61: | ||
|[[Image:open.jpg|thumbnail|350 px|Figure 2b. Closed-Loop configuration (K079020)]] | |[[Image:open.jpg|thumbnail|350 px|Figure 2b. Closed-Loop configuration (K079020)]] | ||
|} | |} | ||
| - | |||
| - | |||
<br> | <br> | ||
Revision as of 02:23, 30 October 2008
| HOME | PROJECT | TEAM | SOFTWARE | MODELING | WET LAB | LAB-BOOK | SUBMITTED PARTS | BIOSAFETY AND PROTOCOLS |
|---|
Contents |
Ecoli.PROM: an Erasable and Programmable Genetic Memory in E. coli
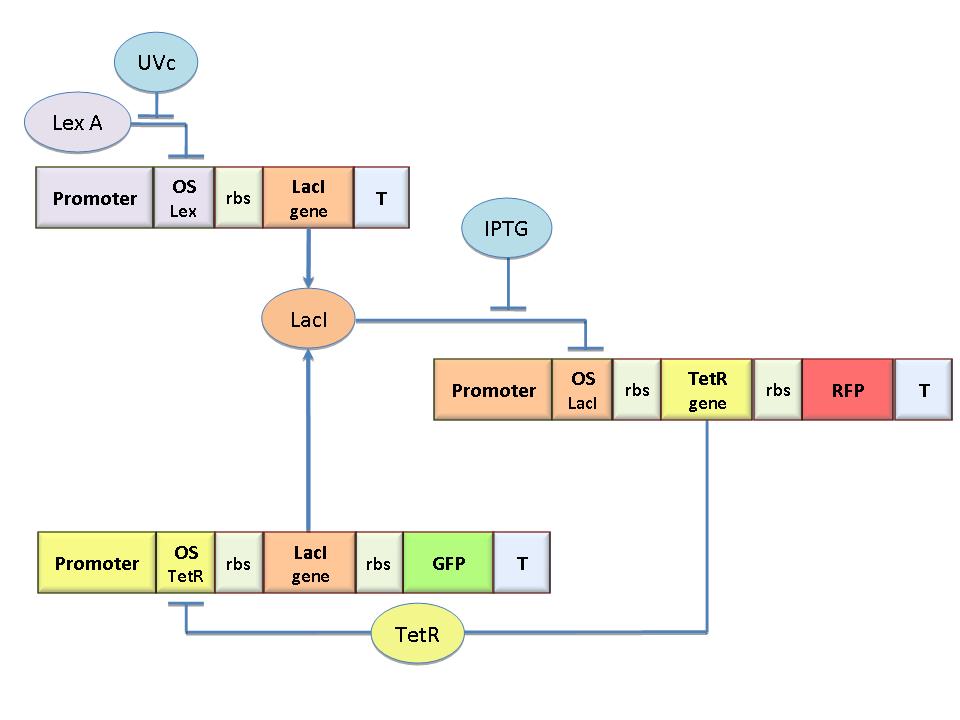
The specific goal of our project was to design a bacterial reprogrammable memory, i.e. colonies of genetically engineered E. coli immobilized in solid medium where they work as an array of binary memory cells. To engineer the bacteria we designed a modular genetic Flip-Flop composed of two parts (Figure 1): a binary memory block and an induction block, sensitive to UVc radiation. UVc has been chosen to have a fine spatial selectivity in programming the memory cells, whereas IPTG should be used to reset the entire memory.
The molecular circuit (see Figure 1) can switch between two different stable states (LacI-ON and TetR-ON), driven by the external stimuli UVc and IPTG. LacI-ON represents the stable state in witch LacI gene is active and LacI protein represses the TetR gene expression, with a positive feedback. Therefore, the LacI-ON state coincides with the TetR-OFF condition. On the contrary, the TetR-ON represents the state with the TetR gene active and the LacI gene silenced (LacI-OFF). Owing to the coexistence of two stable states (bistability), this circuit is capable of serving as a binary memory. We denominated it genetic Flip-Flop since it works as a SR Latch: LacI state is the ![]() output and TetR state is the
output and TetR state is the ![]() output. Uvc is the set signal and IPTG is the reset signal. Indeed, IPTG stimulation inhibits LacI repressor, thus can cause the transition from the LacI-ON state to the TetR-ON. UVc radiation, inactivating LexA repressor through the SOS response [Friedberg et al. 1995] can cause the opposite transition from LacI-ON to TetR-ON.
output. Uvc is the set signal and IPTG is the reset signal. Indeed, IPTG stimulation inhibits LacI repressor, thus can cause the transition from the LacI-ON state to the TetR-ON. UVc radiation, inactivating LexA repressor through the SOS response [Friedberg et al. 1995] can cause the opposite transition from LacI-ON to TetR-ON.
The core elements of this epigenetic memory are the two mutually regulated promoters, each designed as a constitutive promoter flanking an indipendent operator site. In this way, the promoter transcriptional strength and the repressor binding affinity can be independently fixed. Mathematical model analysis and computer simulations were used to obtain a rational design of the regulated promoters. By the model we found the analytic relationships to quote the regulated promoter in terms of transcriptional strength and sensitivity to the repressor and we established such a relevant circuit properties as the bistability and the dynamical response to inputs.
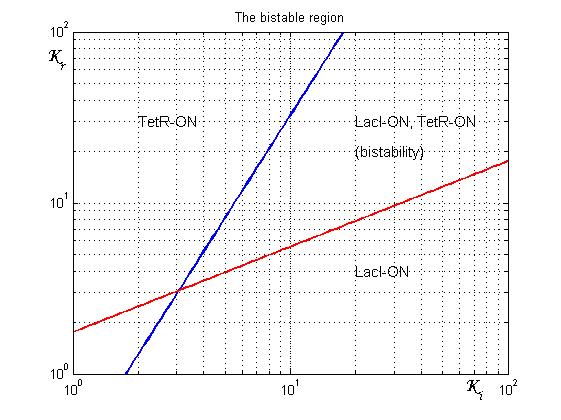
As shown in Figure 2, the coexistence of two stable equilibria (LacI-ON and TerR-ON) is guaranteed for![]() and
and ![]() model parameters greater than 3 in a large range of values (bistability range). Imposing a value for
model parameters greater than 3 in a large range of values (bistability range). Imposing a value for ![]() greater than 3 it is possible to establish a range for
greater than 3 it is possible to establish a range for ![]() to have a bistable behaviour. More the (
to have a bistable behaviour. More the (![]() ,
,![]() ) point is near the bifurcation lines more the transition from bistability to monostability is probably. To have a good robustness we fixed
) point is near the bifurcation lines more the transition from bistability to monostability is probably. To have a good robustness we fixed ![]() and
and ![]() equal to 10. For this value the circuit is sufficently distant from the bifurcation lines to evoid random memory switching but it is possible to set (or to reset) the memory with proper UV and IPTG stimulation.
equal to 10. For this value the circuit is sufficently distant from the bifurcation lines to evoid random memory switching but it is possible to set (or to reset) the memory with proper UV and IPTG stimulation.
Since the Registry lacks promoters with the K parameter equal to 10, we decided to introduce operator site parts in order to build regulated promoter with the fixed transcriptional strenght and repression sensitivity. Thus, we synthesized four libraries of operator sequences, respectively for [http://partsregistry.org/wiki/index.php?title=Part:BBa_K079045 LacI], [http://partsregistry.org/wiki/index.php?title=Part:BBa_K079046 TetR], [http://partsregistry.org/wiki/index.php?title=Part:BBa_K079047 cI] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K079048 LexA] repressor proteins ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K079045 see details]). In each library, there are three sequences each with a different repressor binding affinity to the repressor protein. We isolated each operator with the intention to clone them into BioBrick standard assembly plasmids.
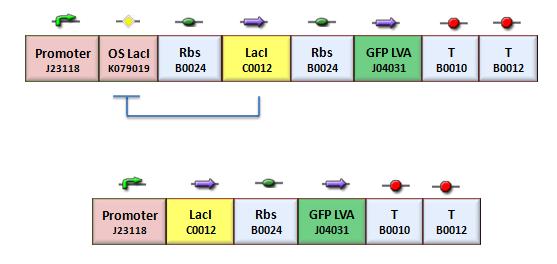
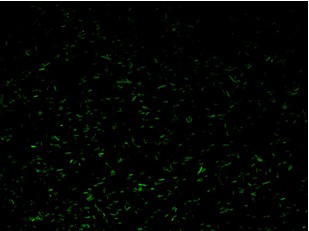
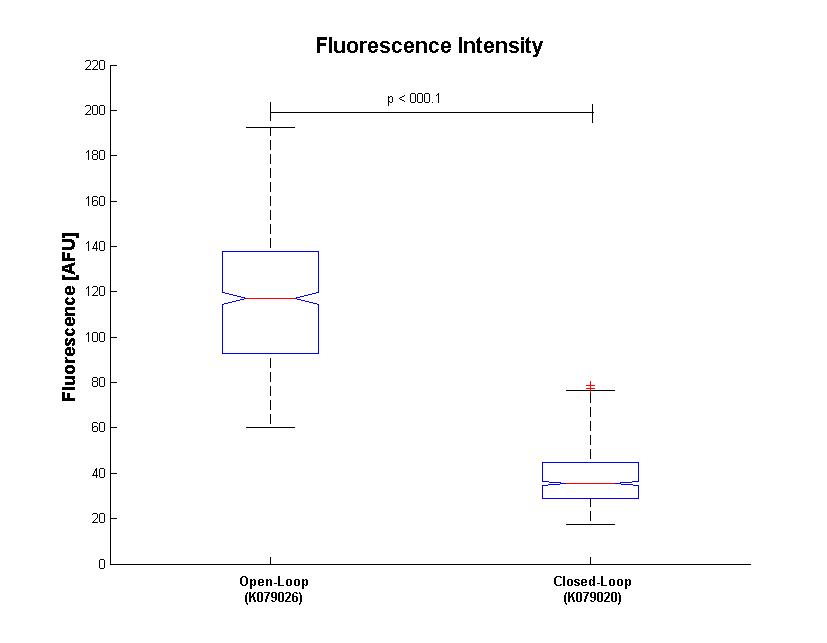
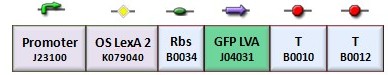
We assess the functionality of independent operators by assembling one circuit where the constitutive promoter BBa_J23118 and LacI operator 2 were auto-regulated by the LacI repressor. GFP was used as the reporter. An identical construct without the LacI operator site was used as the negative control (Figure).
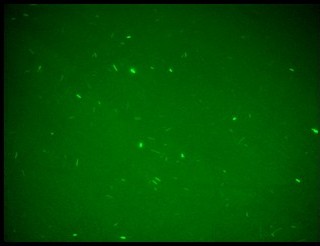
Fluorescence images demonstrate the repression due to the presence of the Lac operator.
In the genetic Flip-Flop, the amount of LacI to set ON the memory is induced by an UV-sensitive trigger . One new construct (K079050) was assembled to test the UV induction. This UV test circuit is presented schematically in Figure 4-5.
|
ConclusionsThis will allow the rational design of regulated promoter elements that are still lacking in the Registry. This approach has been applied in the building of our UV-programmable memory, and we expect it to be a general benefit in a larger number of applications in Synthetic Biology. Collaboration with other iGEM2008 Teams
Concluding the iGEM 2007 ProjectIn the iGEM 2007 we used the LacY gene (BBa_J2210) to design a genetic Schmitt trigger. Since this part was not working well, we sent it to be sequenced and found that it contained a 35 bp insertion upstream the endogenous LacY gene sequence. This insertion probably caused a frameshift in protein translation, making the gene ineffective. So, we amplified the right gene sequence and put it in the BioBrick format. Successive sequencing confirmed the right assembling of this part. We also measured IPTG-induced fluorescence in the genetic Schmitt trigger (see Figure) and we assessed the correct function of the new LacY part. To contribute to Registry’s improvement we decided to send this new part to the Registry ([http://partsregistry.org/Part:BBa_K079015 K0790015]). 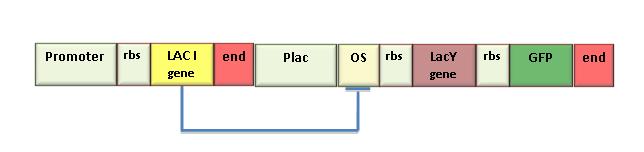 Schematic representation of the genetic Schmitt Trigger. The LacI generator module was included to have a constitutive synthesis of LacI repressor protein witch makes up for endogenous LacI. LacY permease introduces a positive feedback. GFP is the reporter for pLac activation. This circuit express high level of fluorescence with very low concentration of inducer (IPTG=1uM). After switching, the fluorescence level f is insensitive to an increase in inducer dosage.
|
 "
"