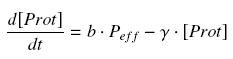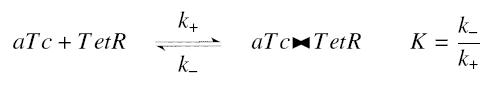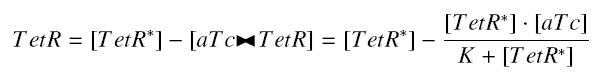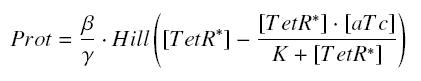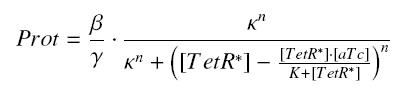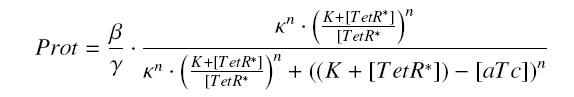Team:Paris/Modeling/estimation
From 2008.igem.org
(→what are we looking for ?) |
|||
| Line 44: | Line 44: | ||
===how to control the concentration of the transcription factor ?=== | ===how to control the concentration of the transcription factor ?=== | ||
| - | ====By aTc-TetR-''Ptet''==== | + | <center>====By aTc-TetR-''Ptet''====</center> |
Now, we must use as a variable of reference an element that could be introduced in the bacteria, well-controlled, and from which will depends all the concentrations of our transcription factor. We propose a construction in which our transcription factor is put after the promoter ''Ptet'', which is under the repression of TetR. Since aTc is a small diffusive molecule that binds to TetR and inhibits this way the repression of ''Ptet'', we can use it as an 'inducer'. To do so, we must place in the bacterium the gene ''tetR'' after a constitutive promoter (like J23101). According to previous hypothesis, this will provide at steady-state a 'constant concentration' of TetR (we note [TetR*], and it is supposed to be the TOTAL concentration of TetR, under every form) in the bacterium. If we consider the binding reaction this way (where LacI_IPTG denotes the complex) | Now, we must use as a variable of reference an element that could be introduced in the bacteria, well-controlled, and from which will depends all the concentrations of our transcription factor. We propose a construction in which our transcription factor is put after the promoter ''Ptet'', which is under the repression of TetR. Since aTc is a small diffusive molecule that binds to TetR and inhibits this way the repression of ''Ptet'', we can use it as an 'inducer'. To do so, we must place in the bacterium the gene ''tetR'' after a constitutive promoter (like J23101). According to previous hypothesis, this will provide at steady-state a 'constant concentration' of TetR (we note [TetR*], and it is supposed to be the TOTAL concentration of TetR, under every form) in the bacterium. If we consider the binding reaction this way (where LacI_IPTG denotes the complex) | ||
| Line 78: | Line 78: | ||
Then, it is still interesting to compare the three possibilities, if we have enough time... | Then, it is still interesting to compare the three possibilities, if we have enough time... | ||
| - | ====By AraC-Arabinose-''Pbad''==== | + | <center>====By AraC-Arabinose-''Pbad''====</center> |
An other well-known promoter called ''Pbad'', is induced by the complex AraC_-_Arabinose, where AraC appears to be a protein ''constitutively produced'' by an operon attached to ''Pbad'', and Arabinose is a sugar, that we can add at will in the medium, and diffuses through cells. | An other well-known promoter called ''Pbad'', is induced by the complex AraC_-_Arabinose, where AraC appears to be a protein ''constitutively produced'' by an operon attached to ''Pbad'', and Arabinose is a sugar, that we can add at will in the medium, and diffuses through cells. | ||
Then, our "inducer" will be the Arabinose, that we will treat as an ''abstract variable'' without intending to know the concentrations of AraC, or of the complex. | Then, our "inducer" will be the Arabinose, that we will treat as an ''abstract variable'' without intending to know the concentrations of AraC, or of the complex. | ||
| + | |||
| + | ===The Problem of the RBS=== | ||
| + | |||
| + | The promoter is not the only one factor which control the expression of the protein... in particular, the ''traduction phenomenon'' is almost as important as the ''transduction''. As we decided not to take into acount the traduction, that means that we do not want to deal with mRNA and ribosomes ; nevertheless, as we aimed in the "hill approach" to simulate as precisely as possible the involved concentrations, we must ''integrate in the traduction simulation the most important influences of the transduction'''. | ||
| + | |||
| + | In particular, since the ''traduction rate'' depends near linearly (as we guess ; it gives the affinity between the mRNA and the ribosome !) of the '''Ribosome Binding Site''', it can be modelised in the '''beta coefficient of the promoter''' (see [[Team:Paris/Modeling/Programs#Finding_Parameters]]) (even if, rigorously, we must describe such induction by the association "promoter + specific RBS"). | ||
| + | |||
| + | Still, as it is proposed by iGEM, we will use the GFPgenerator (E0240) in association with its RBS (E0032), to caracterise the '''expression of the gene behind promoters'''. However, the RBS before the genes which codes for the ''transcriptions factors'' we want to induce, are the natural RBS (specific respectively to ''tetR'', ''flhDC'', ''fliA'', etc... ). Therefore, we must pay attention on what we are measuring. | ||
===what are we looking for ?=== | ===what are we looking for ?=== | ||
Revision as of 14:19, 16 August 2008
|
 "
"