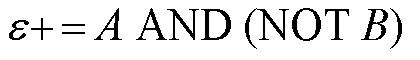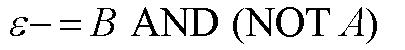Team:Minnesota/Project
From 2008.igem.org
(→Project Details) |
(→Project Details) |
||
| Line 50: | Line 50: | ||
|[[Image:BasicComparatorMN.jpg|thumb|500px|center|Basic Function of Comparator | |[[Image:BasicComparatorMN.jpg|thumb|500px|center|Basic Function of Comparator | ||
]] | ]] | ||
| + | |- | ||
| + | | | ||
|- | |- | ||
|<div align="center"><gallery perRow="2" widths="300px" heights="200px"> | |<div align="center"><gallery perRow="2" widths="300px" heights="200px"> | ||
Revision as of 16:40, 26 June 2008
| The Minnesota iGEM '08 team consists of groups working on two different projects. One team is located primarily in Minneapolis and is designing a comparator for their project. The other team is based in St. Paul and is working on a time bomb. | |
|
| |
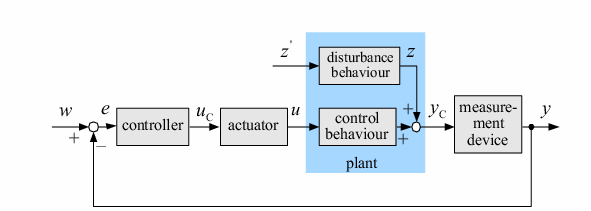
| Control systems are an integral component of almost all aspects of life. Whether it is in industrial, biological, or chemical applications, controllers provide a way to keep systems functioning properly. A vital part of any control system is the comparator. This component compares a set point value and a measured value, and determines which is larger. It then sends the appropriate signal to the controller, which reacts to bring the system back to the set point. In typical applications, the controller equipment is electronic. However, our team set out to create a comparator using only genetic components. This comparator could potentially be used as part of a new, solely biological control system that could be used to treat many diseases afflicting humans, for example diabetes. This comparator could compare the blood sugar of a patient to what it should be, and send this result to a control system that could compensate, for example by changing the levels of insulin.
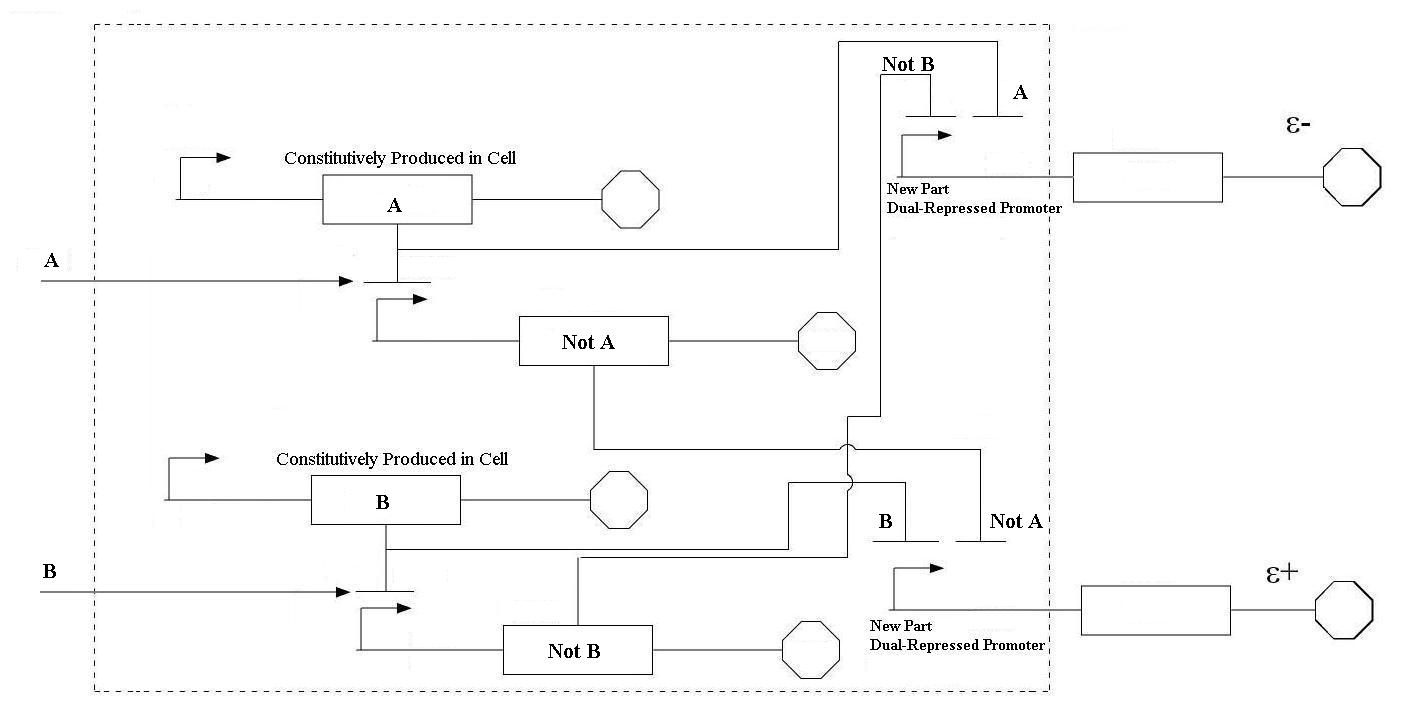
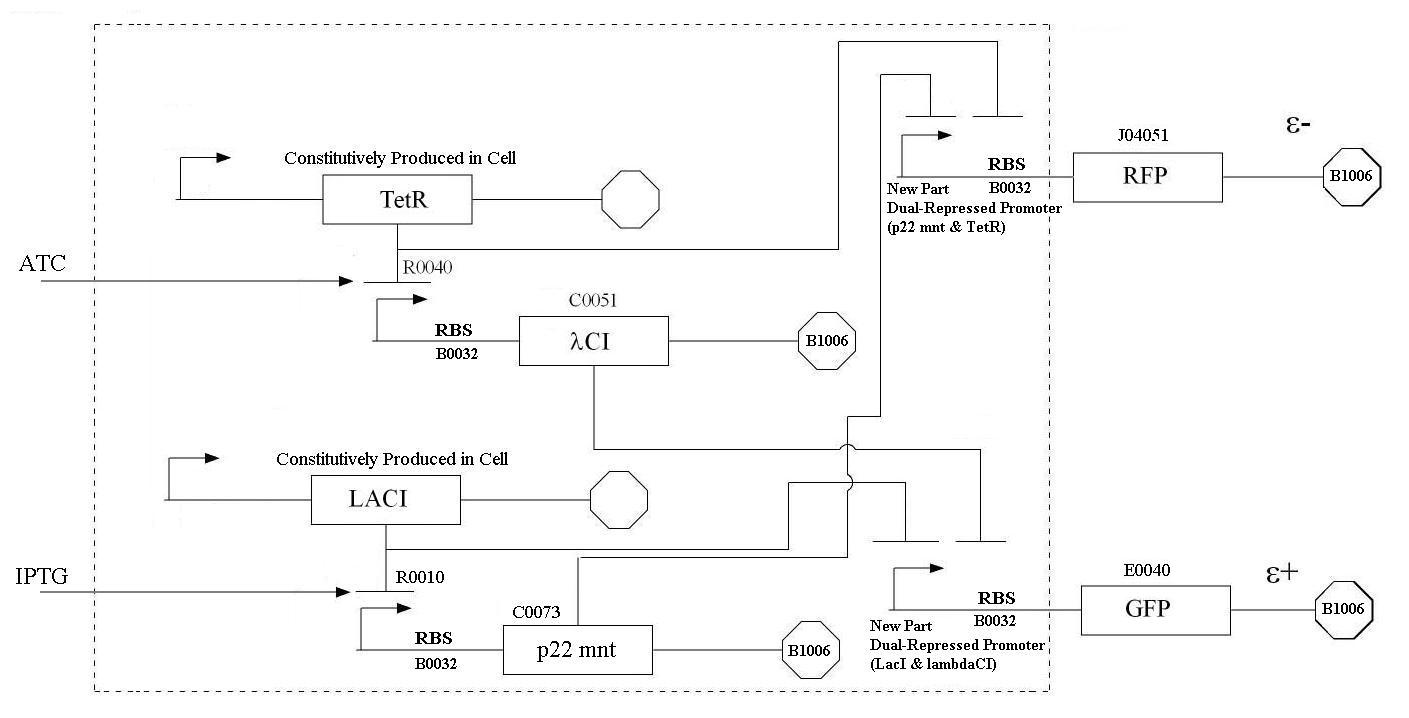
In order to undertake this task, a system involving six genes was designed. For our system, the two inputs (one representing the set point and one representing the measured value) are IPTG and ATC. These inputs will activate the transcription of the LacI and TetR proteins, and set in motion the rest of the system to produce the outputs. Depending on the amounts of the two inducer molecules added to the system, either green fluorescent protein (GFP) or red fluorescent protein(RFP) will be produced. The actual design of the system can be seen below. In order to complete this project, a total of six genes will have to be cloned into plasmids, and two new BioBrick parts will be produced. One will be a TetR and p22-MNT dual-repressed promoter, and the other will be a LacI and lambdaphage cI dual-repressed promoter. We will also build mathematical models and conduct computer simulations that will help with the designs. This project will pave the way for other parts of a true genetic PID controller to be produced, which could be an exciting scientific development in the near future.
| |
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Contents |
Overall project
Control systems are an integral component of almost all aspects of life. Whether it is in industrial, biological, or chemical applications, controllers provide a way to keep systems functioning properly. A vital part of any control system is the comparator. This component compares a set point value and a measured value, and determines which is larger. It then sends the appropriate signal to the rest of the system. In typical applications, this system is electronic. However, our team set out to create a comparator using only genetic components. This comparator could potentially be used as part of a new, solely biological control system that could be used to treat many diseases afflicting humans, for example diabetes. This comparator could compare a diabetics blood sugar to what it should be, and send this result to a control system that could compensate.
In order to undertake this task, a system involving six genes was designed. For our system, the two inputs (one representing the set point and one representing the measured value) are IPTG and ATC. These inputs will activate the transcription of the LacI and TetR proteins, and set in motion the rest of the system to produce the outputs. Depending on the amounts of the two inducer molecules added to the system, either green fluorescent protein(GFP) or red fluorescent protein(RFP) will be produced. The actual design of the system can be seen below. In order to complete this project, a total of six genes will have to be cloned into plasmids, and two new BioBrick parts will be produced. One will be a TetR and p22 mnt dual-repressed promoter, and the other will be a LacI and lambdaphage cI dual-repressed promoter. This project will pave the way for other parts of a true genetic PID controller to be produced, which could be an exciting scientific development in the near future.
Project Details
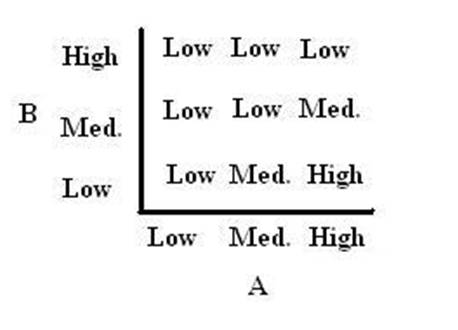
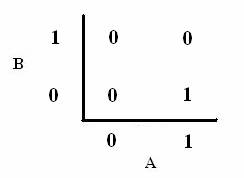
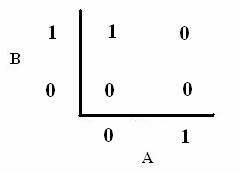
| In order to design a genetic comparator, the team first looked at an electronic comparator to try to emulate it. A typical comparator has two inputs: A is the set point value, which is input into the system, and B is the measured value, which is found by measuring the controlled variable and sending this result back to the comparator. Once the comparator recieves these two values, it compares them and produces a signal. Typical this signal is positive or negative. However, for a genetic controller, there is no such thing as a negative growth, so the genetic system was forced to have two outputs, instead of the one found on typical systems. These outputs were called epsilon+ and epsilon- to distinguish whether the signal was positive or negative. A black-box comparator can be seen below. |
 "
"