Team:Heidelberg/Project/Killing I
From 2008.igem.org


Phages - Project Description
Antibiotics are a very powerful way to cure bacterial infections till today, but their power withers. More and more bacterial strains become resistant against different antibiotics, due to the even greater powers of evolution: chromosomal mutations do not only make them resistant against natural antibiotics, but also against newly developed synthetic ones [1]. Bacteria also have a very effective way to share those resistances with each other, not limited by species boundaries: the transfer of plasmids by conjugation (which will be investigated in more detail in the second part of the project). Multiresistant bacterial strains like Pseudomonas aeruginosa, Staphylococcus aureus and Enterococcus faecalis are a serious problem for the usage of antibiotics aginst infectious deseases, because they can give their plasmids to other pathogens and cannot be killed by ordinary antibiotics.
Nevertheless, there are some alternatives to antibiotics, which are currently researched: the usage of bacterial toxins, the usage of probiotic bacteria and the usage of bacteriophages as antibiotics. In our project we focus on two approaches: To engineer a probiotic bacteria which uses toxins as a killing device and one which uses bacteriophages, respectively.
Our part of the team is concerned with the phage strategy. Bacteriophages are a group of viruses, for which bacteria are the natural hosts, among them human pathogens. There are two types of phages: virulent and temperent ones. The virulent ones infect a host cell and multiply by using the host resources and lyse the host to be set free. The temperent ones are able to either integrate into the host chromosome (lysogenic) or follow a lytic strategy and multiply and thereby lyse the cell. Virulent, non-lysogenic phages are used for the treatment of infectious diseases caused by bacteria since the 1930s. Phage therapy was intensely used since the first antibiotics where discovered in the 1940s. Later, this technique was extensively investigated and used especially in the Soviet Union [2]. Since there are more and more multiresistant bacterial strains, phage therapy becomes interesting again [3].
The special quality of our approach is that we use bacteria as vectors for the phages. This will avoid the consequences of a systemic application of bacteriophages, because they are only released at the site of infection. Smaller amounts of phages are needed, because they are not spreading through the whole body.
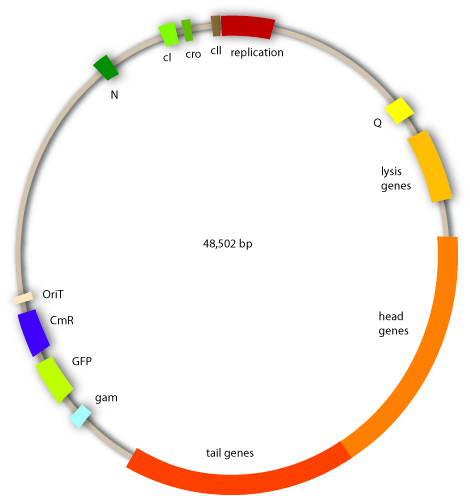
The bacteriophages we use are lambda phages, because this project should be a proof of principle for our technique, and therefore we rely on this very well investigated type of phage. The lambda phages are a class of temperent bacteriophages. They have a 48,502 bp genome consisting of linear double stranded DNA. The genome is a highly mosaic, modular structure. Like all viruses they carry out distinct steps of phage infection. The first step is adsorption. Lambda phages have receptors at their tail fibres, which bind specifically to mannose transporters of E.coli. These receptors are not able to bind stably to structures on other bacteria strains and are therefore responsible for the host specificity of the lambda phage. After adsorption, the lambda phage injects its genome into the host cell. In the environment of the host, the linear DNA circularizes at the so called cos sites: these are 12bp complementary overhangs at the ends of the genome.
As temperent phages the lambda phages are able to undergo either a lytic or a lysogenic cycle. Undergoing a lysogenic cycle, the infecting phage integrates its genome into the host genome and stays there as a stable prophage. In this state it will be replicated passively with the host genome. It will not lyse the cell and multiply without an extern trigger – for example if the host cell is stressed by DNA damage. If carrying out a lytic cycle, the lambda phage also injects its genome into the host cell. This will then be replicated autonomously, viral proteins will be produced, first for the formation of new phage particles, which are formed via self assembly, and second for the lysis of the host cell, upon which the newly produced phages are released. The lambda phage has a very complex regulatory network that controls which cycle to carry out.
The default mode of this network is the lytic growth. This is carried out in three steps: 1. After the injection of the genome the transcription starts from the promoters pL and pR. This early transcription has two products - the regulators N and cro - and is attenuated at the terminators tL1 and tR1. Cro acts as a weak repressor for the pL and pR promoters, allows little but steady transcription from these promoters and thereby facilitates the lytic state (because cII concentrations are kept low – see below). N is an antiterminator that enables the RNA polymerase to overcome the tL1 and tR1 terminators and to read the DNA up to the sib and the tR2 terminator, respectively. 2. With the help of N, the delayed early genes are transcribed. The products are the lysogenic regulators cII and cIII as well as the lytic DNA replication functions O and P and the late gene regulator Q. (N is critical for the switch to the lytic state: cells carrying N on a plasmid and thereby expressing it strongly allow a very effective lytic mode of the phage – they from clear plaques!) 3. An accumulation of Q makes the RNA polymerase resistant to the tR’ terminator and thereby allows expression of the late genes, which are essential for the lytic cycle. They encode for the viral coat proteins and for lysis proteins. These late genes are expressed with a delay after the delayed early genes, which results from the threshold concentration of Q that has to be accumulated to overcome the tR’ terminator.
In contrast, the lysogenic state is also initiated during the transcription of the early delayed genes: 1. Under lysogenic conditions, cII is able to accumulate over a threshold level. Then it can bind to the pI, pRM and paQ promoters, thereby initiating the expression of int, cI and repressing the expression of Q, respectively. The int gene encodes a protein called integrase, which integrated the phage genome into the host genome. 2. CI binds to the early operators oL and oR and represses transcription from pL and pR. Repression of transcription from pR inhibits the transcriptional activation of the ori of the lambda phage genome and thereby replication. So cI maintains the stable lysogenic state after cIi has switched the genetic cascade of the lambda phage in favour of lysogeny. 3. During the prophage state, cI is expressed continuously. If another phage injects its genome into the host cell of a prophage, cI binds to regulatory sequences on this second genome and prevents any transcription, thereby making the host immune to super infections.
The decision for the lytic or lysogenic mode depends on the multiplicity of infection and the physiological conditions of the bacteria cell. Many phages infecting a cell favour lysogeny. Rich nutrition of the bacterium favours the lytic state, because it increases N expression levels. Since lytic and lysogenic genes are under the control of the same promoters, the critical point for the decision of a state are the RNA and protein levels, influenced by the activity of degrading enzymes [4][5][6][7][8][9].
Our work focussed on designing a non-lysogenic phage, because we wanted to produce an efficient killing module for the probiotic bacterium. In preventing the phage from undergoing a lysogenic cycle, we assure that it will kill the target bacteria as fast as possible and without extern triggers. For this task we aimed to discard the int gene from the phage genome, so that it would be incapable of integrating into the host genome and being lytically inactive.
Cloning work with the lambda phage genome is tricky, because it contains few single cutter restriction sites. Therefore we had to cut out one fragment containing the int, the xis and another gene, gam. Xis and int can and shall be discarded, but the gam gene codes for a protein that prevents the overhanging single strains at the cos sites from being degraded by host enzymes. So we would need the gam gene in our engineered phage. This is why we designed an insert for the phage to put in the remaining genome after cutting out the fragment containing xis, int and gam. This insert contains gam, and an antibiotic selection marker to be able to select those bacterial clones that contain the lambda genome.
We also added a GFP gene for visualizing and measurement purposes. To be able to later modify this phage insert again, we added special restriction sites to only cut out the non-phage genes.
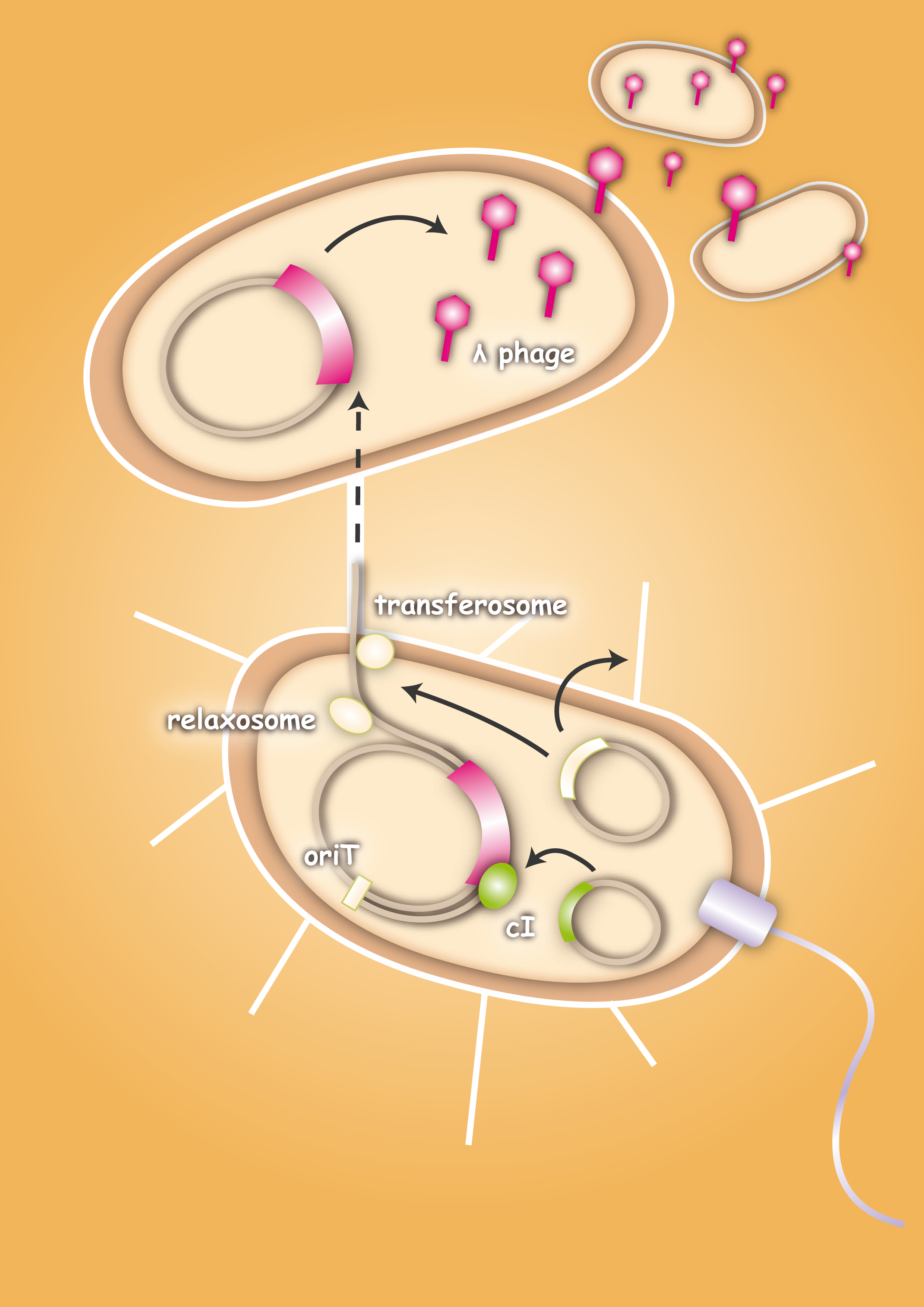
When designing the probiotic bacterium to kill pathogens, we had one problem to solve: the probiotic bacteria should not be lysed by the release of the phages, but the phages had to get into the target bacteria. Therefore we decided to use conjugation for this task. Conjugation is the most important bacterial tool to transfer genetic information between and across species. It enables them to adapt very fast to environmental conditions - antibiotic resistances are also spread via conjugation. The clou: our technique was developed to fight infectious diseases which cannot be treated with antibiotics any more because of these resistances. Therefore we try to overcome the pathogens with their own weapons.
In general, conjugation is the unidirectional transfer of genetic information from a donor cell to a recipient. This genetic information can be a conjugative or mobilizable plasmid or a transposon. Normally, conjugation needs direct cell contact and the formation of a plasma bridge between the cells.
A conjugative plasmid of gram negative bacteria contains a characteristic set of genes and sequences. The tra locus includes the genes for the sex pili and for proteins that carry out synthesis, replication and transfer of the DNA during mating.
The sex pili are formed by pilin and regulatory proteins. These are polymeric structures that extend into the periphery from the surface of the donor cell and can attach to the surface of other bacteria and initiate conjugation. They do make the physical contact between the cells by retracting, but they do not form the channel, through which the DNA is shuttled.
After the initiation of conjugation, a relaxase creates a nick in one of the strands of the conjugation plasmid at the nic sequence of the origin of transfer (oriT) and unwinds the strand to be transferred (the T-strand). If the relaxase (TraI in IncP plasmids) works in complex of many proteins, this is called the relaxosome. The T-strand is unwound from the plasmid and transferred into the recipient in a 5’-to 3’ terminus direction. During this process, the relaxase protein stays bound to the 5’ end of the T-strand. The transfer is catalysed by an enzyme (called TraG in IncP plasmids), which possibly acts by connecting the relaxosome at the 5’ end of the T-strand with the mating complex (mating complex are all proteins that are important for the formation of the plasma bridge between the two cells). The remaining strand is replicated using rolling circle replication. The transferred strand is complemented in the recipient cell.
Since a conjugation plasmid contains all the information necessary to transfer DNA, the recipient becomes a donor after conjugation [10].
There exist many variations of this classic system of conjugation. One is a two component system, consisting of a helper plasmid and a mobilizable plasmid. Most of the proteins needed for conjugation are trans acting elements, they do not have to be on the transferred plasmid, but can either be part of the chromosome or be on another plasmid, a so called helper plasmid. The only cis acting element needed for the transfer of a plasmid is the oriT sequence. It consists of the sequence, where the relaxase nicks the DNA, the nic, and some genes called mobilizable (mob) genes. Together they make about 500bp. The mob genes encode for example accessory proteins that bind to the DNA sequence at and around the nic sequence and assist the relaxase. If they are included in an arbitrary plasmid, it will be mobilized and conjugated by the helper plasmid. This two component system has some important advantages: You do not need to put the genetic information you want to transfer on a big conjugation plasmid like an F-plasmid or an RP-plasmid. This makes conjugation much faster and more effective, because the time and the effectivness of conjugation depend on the length of the plasmid to transfer. Additionally, you can use every kind of plasmid as a mobilizable plasmid. This makes it perfect for our purpose: We can use the lambda genome as a mobilizable plasmid in cloning an oriT into it and providing a suitable helper plasmid.
There are different groups of conjugative plasmids that can also function as helper plasmids in enterobacteria: the classic F-plasmid, RP4/RP1 among others. We decided to use an IncP (incompatibility group P) plasmid and a complementary oriT as our conjugation system, because they proofed to be very efficient [11]. These are plasmids with a broad host range that carry multiple antibiotic resistance genes. The plasmids of this group do not need host-chromosome encoded components for the mobilization of plasmids, like RSF1010 for example needs [12]. Therefore they have a very broad range of hosts and fit in very well in the modular concept of synthetic biology.
The prototype of InP plasmids is the RP4 plasmid, which is a close relative of the RP1 plasmid of which we use a derivative. The tra genes are organized in two operons, tra1 and tra2. Tra1 encodes for all of the DNA transfer and replication system, especially for the relaxosome, the complex, which initiates the transfer replication [14]. Tra2 encodes for proteins necessary for mating pair formation.
The plasmid we use is called pUB307 [15][16]. It is a derivate of RP1, which lacks the transposon Tn1 (Ampicillin resistance cassette) by spontaneous excision.
To avoid that our probiotic bacteria, which would contain and transfer the phage to the target pathogenic bacteria, would also be killed by the phage, we used the natural way to make them immune. Therefore we took the lambda cI gene and put it behind a constitutive promoter. The cI protein will then prevent the expression of the lambda genome in our bacteria and prevent them from being lysed.
Outlook
The conjugation system may allow a broader application of our system in terms of species. Since the RP-plasmids are broad host range plasmids, and can therefore transfer genetic information to a variety of gram negative and gram positive bacteria species. There is also evidence for conjugative transfer between bacteria and yeast cells [17]. If one could use this system to transfer genetic information into eukaryotic cells, this could be an application to cure plant diseases or even human diseases by injecting for example antisense RNA to silence viral replication or oncogenes.
References
[1]Cirz RT, Chin JK, Andes DR, de Crécy-Lagard V, Craig WA, Romesberg FE (2005). "Inhibition of mutation and combating the evolution of antibiotic resistance". PLoS Biol. 3 (6): e176
[2]N Chanishvili, T Chanishvili, M. Tediashvili, P.A. Barrow (2001). "Phages and their application against drug-resistant bacteria". J. Chem. Technol. Biotechnol.) 76: 689–699
[3]Weber-Dabrowska B, Mulczyk M, Górski A (Jun 2003). "Bacteriophages as an efficient therapy for antibiotic-resistant septicemia in man". Transplant. Proc. 35 (4): 1385–6
[4]SWITCHES IN BACTERIOPHAGE LAMBDA DEVELOPMENT
Amos B. Oppenheim, Oren Kobiler, Joel Stavans, Donald L. Court, Sankar Adhya
Annual Review of Genetics, December 2005, Vol. 39, Pages 409-429
[5]Brussow, H. and Hendrix, R.W. 2002 "Small is beautiful.", Cell, 108, 13-16
[6]Dodd, J.B. Shearwin, K.E. and Egan, J.B. 2005 "Revisited gene regulation in phage lambda.", Curr Opin Genet Dev, 15, 145-152
[7]Friedman, D.I. and Court, D.L. 2001 "Bacteriophage lambda; alive and well and still doing its thing.", Current Opinion in Microbiology, 4, 201-207
[8]Court, D.L. et al. (2007): A new look at bacteriophage lambda genetic networks. In: J. Bacteriol. Bd. 189, 298-304
[9]Gottesman, M.E. & Weisberg, R.A. (2004): Little lambda, who made thee? In: Microbiol. Mol. Biol. Rev. Bd. 68, S. 796-813
[10]Holmes RK, Jobling MG (1996). Genetics: Conjugation. in: Baron's Medical Microbiology (Baron S et al, eds.), 4th ed., Univ of Texas Medical Branch
[11]Introduction of DNA into Actinomycetes by bacterial conjugation from E. coli—An evaluation of various transfer systems, F. Blaesing, A. Mühlenweg, S. Vierling, G. Ziegelin, S. Pelzer, E. Lanka, Journal of Biotechnology 120 (2005) 146–161
[12]Lanka E, Wilkins BN (1995) DNA processing reactions in bacterial conjugation. Annu Rev Biochem 64:141-169
[13]Complete nucleotide sequence of Birmingham IncP a plasmids, E. Lanka et a., J. Mol. Biol., (1994), 239, 623-663
[14]Lessl M, Balzer D, Weyrauch K, Lanka E (1993) The mating pair formation system of plasmid RP4 desined by RSF1010 mobilization and donor speci®c phage propagation. J Bacteriol 175:6415-6425
[15]Target Choice and Orientation Preference of the Insertion Sequence IS903
Wen-Yuan Hu and Keith M. Derbyshire, J Bacteriol, June 1998, p. 3039-3048, Vol. 180, No. 12
[16]Transposition of TnA Does Not Generate Deletions, Peter M. Bennett, John Grinsted, Mark H. Richmond, Molec. gen. Genet. 154, 205-211 (1977)
[17]Lessl M, Lanka E (1994) Common mechanisms in bacterial conjugation and Ti-mediated T-DNA transfer to plant cells. Cell 77:1-20
 "
"