Team:KULeuven/Model/Filter
From 2008.igem.org
Contents |
Filter
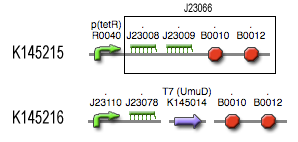
Position in the system
The filter is positioned immediately after the input, because its job is to filter out possible noise signals or background signals that aren't caused by the "desease". It is the starting piece of the whole system, situated before the invertimer- and the reset-subsystem.
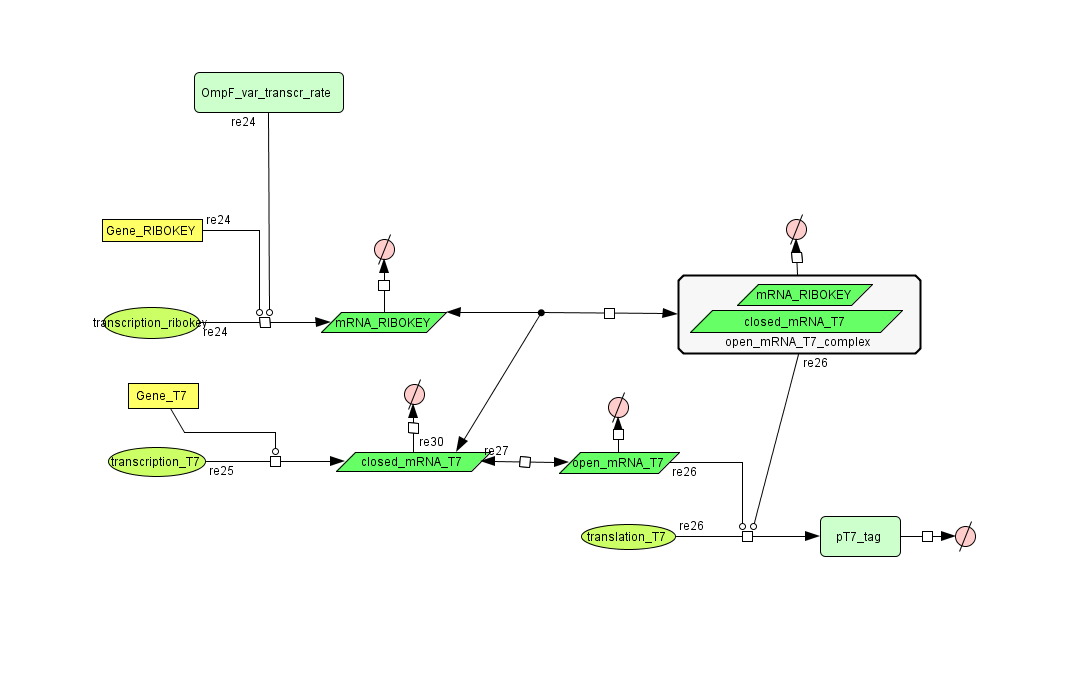
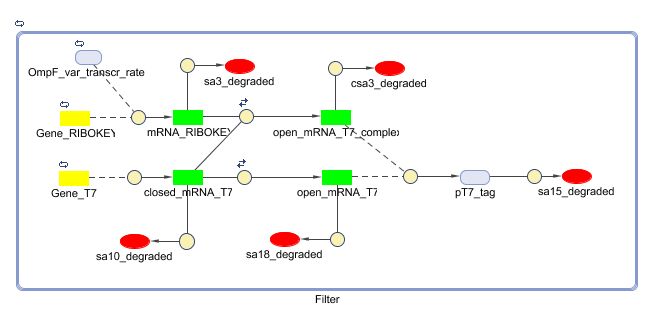
Describing the system
see also: Project:Filter
ODE's
Parameters
| Name | Value | Comments | Reference |
|---|---|---|---|
| Degradation rates | |||
| dpT7 tag | 0.00155 s-1 | UmuD tag added to speed up degradation of otherwise too stable T7 polymerase | [http://www.openwetware.org/wiki/IGEM:Tsinghua/2007/Projects/RAP#Model_and_simulation link] |
| dmRNA RIBOKEY | 0.00462 s-1 | estimate: because this RNA isn't translated, it degrades faster | [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=124983&blobtype=pdf link] |
| dclosed mRNA T7 | 0.00462 s-1 | estimate: because this mRNA isn't translated, it degrades faster | [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=124983&blobtype=pdf link] |
| dopen mRNA T7 | 0.00231 s-1 | [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=124983&blobtype=pdf link] | |
| dopen mRNA T7 complex | 0.00231 s-1 | [http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=124983&blobtype=pdf link] | |
| Equilibrium constants | |||
| Keq 1 | 0,015 [M] | between closed and open T7 mRNA, experimental | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| Keq 2 | 0.0212 [M] | between closed T7 mRNA and key unlocked mRNA complex, experimental | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| Rate constants | |||
| kdis | 0.00416 s-1 | derived from experimental values | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| kcomplex | 0.00237 s-1 | derived from experimental values | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| kclosed | 500 s-1 | derived from experimental values | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| kopen | 7.5 s-1 | derived from experimental values | [http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain link] |
| ktranslation | 0.167 s-1 | lock defined translation rate for T7 RNA pol | [http://partsregistry.org/Part:BBa_B0032 link] |
| Transcription rates | |||
| TetR_var_transcr_rate | p(TetR) dependent | (RiboKey) between 5E-5 and 0.0125 s-1 ~ [aTc] | |
| kmRNA T7 | 0,0011 s-1 | weak constitutive promoter J23109 | [http://partsregistry.org/Part:BBa_J23109 link] |
Remark: The key-lock system has been enhanced to 0.3%-14% (new parameters have been added)
Models
CellDesigner (SBML file)
Matlab (SBML file)
Simulations
1. AND gate of the filter
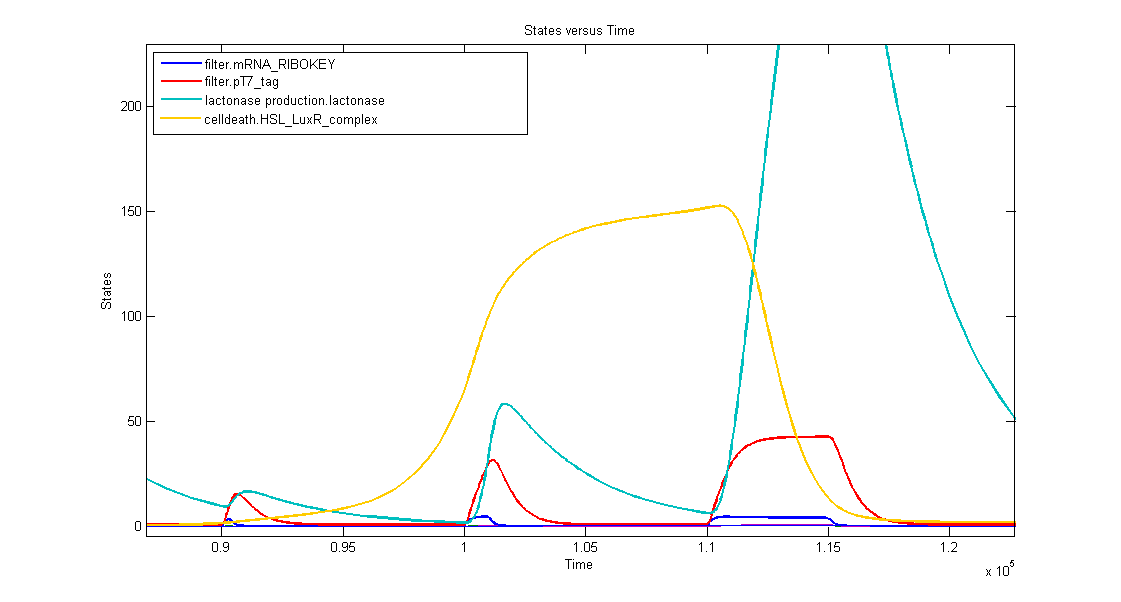
In the simulation we can clearly see this series of events:
- when dark blue(ribokey) starts to increase, red (T7) also starts to increase, giving rise to an increasing amount of lactonase (blue) = AND-GATE.
- when dark blue(ribokey) starts to decrease, red (T7) also starts to decrease, but much slower. The lactonase also starts to decrease, as it should be.
The short lifetime of the ribokey compared to the lifetime of the T7-protein, guarantees that the AND-GATE always works perfectly fine: when there's no more input, the ribokey will rapidly decrease (and disappear) and makes sure that the AND-GATE is not activated anymore, even when the T7-protein is slowly decreasing.
2. Filtering in practice
In the simulation three kinds of inputpulses have been used:
- First pulse: 300s
- small peak of lactonase
- no influence on the timer
- Second pulse: 1000s
- medium peak of lactonase
- influences the timer by levelling the timing capabilities, but it doesn't reset the timer
- Third pulse: 5000s
- huge peak of lactonase
- reset of the timer: amount of complex goes to zero
 "
"