Team:KULeuven/Model/Reset
From 2008.igem.org
Contents |
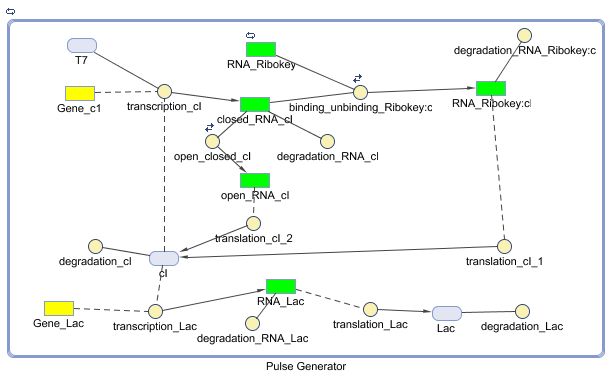
Pulse Generator
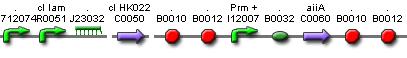
Position in the system
The Pulse Generator-subsystem is directly linked to the Filter.
When the filter indicates that the input is zero (there is no desease), the system will (ideally) produce no lactonase. As soon as the output of the filter is one, the subsystem will produce a pulse of lactonase which will be high enough to 'remove' all HSL present in the system and in that way reset the timer.
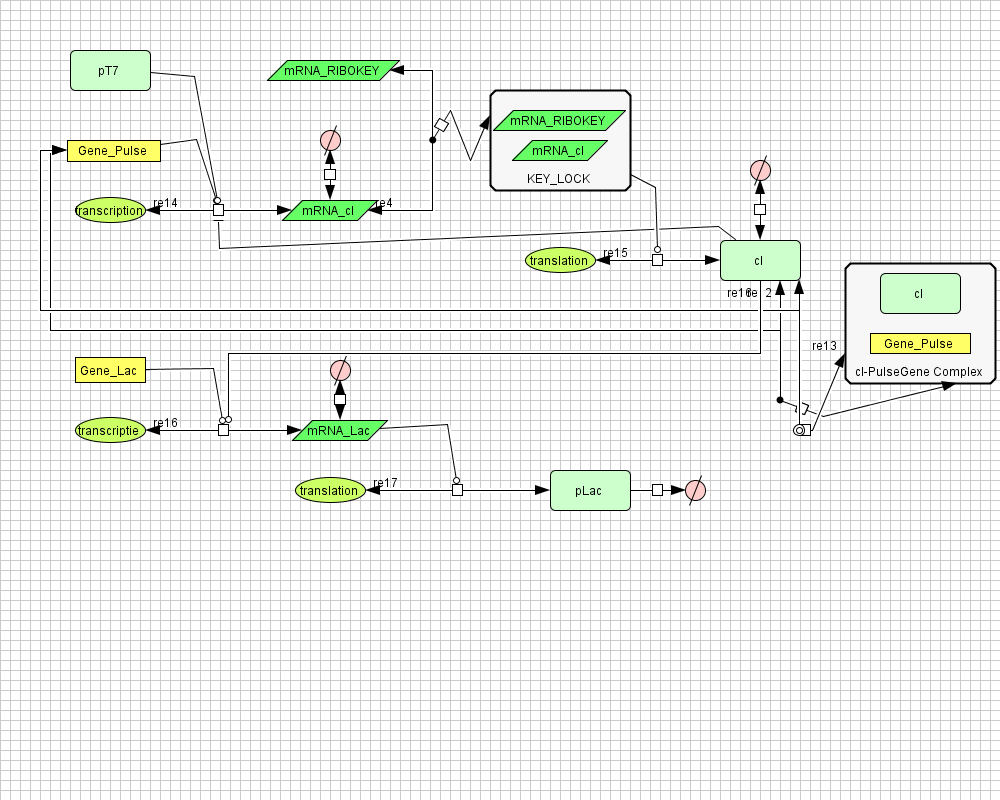
Describing the system
ODE's
![]() NOT AVAILABLE
NOT AVAILABLE
Parameters
| Name | Value | Comments | Reference |
|---|---|---|---|
| Degradation rates | |||
| dRNA_cI | 0.00462 s-1 | ||
| dcI | 7.0E-4 s-1 | [http://parts.mit.edu/igem07/index.php?title=ETHZ/Parameters link] | |
| dRNA_Lac | 0.00231 s-1 | ||
| dLac | 2.888E-4 s-1 | ||
| dRNA_Ribokey:cI | 0.00231 s-1 | ||
| Dissociation constants | |||
| KRibokey:cI | 0.00212 | kass/kdiss for the Ribokey cI complex | |
| KcI | 0.00337 | binding cI on cI-Promotor | [http://parts.mit.edu/igem07/index.php?title=ETHZ/Parameters link] |
| Transcription rates | |||
| kRNA_cI | 0.025 s-1 | maximal transcription rate RNA cI (no cI repressor present) | |
| kRNA_Lac | 0.025 s-1 | ||
| Translation rates | |||
| kcI | 0.167 s-1 | ||
| kLac | 0.167 s-1 | RBS is B0032 (efficiency 0.3) | [http://partsregistry.org/Part:BBa_B0032 link] |
| Hill cooperativity | |||
| ncI | 2.0 | [http://parts.mit.edu/igem07/index.php?title=ETHZ/Parameters link] | |
Models
CellDesigner (SBML file)
Matlab
Problem
The idea of a pulsgenerator as reset mechanism doesn't meet the black-box requirements for the following reasons:
- it takes too long before the proposed system generates a pulse-like event
- the pulse itself is too long
- a constant lactonase production sequence generates enough lactonase to reset the timer
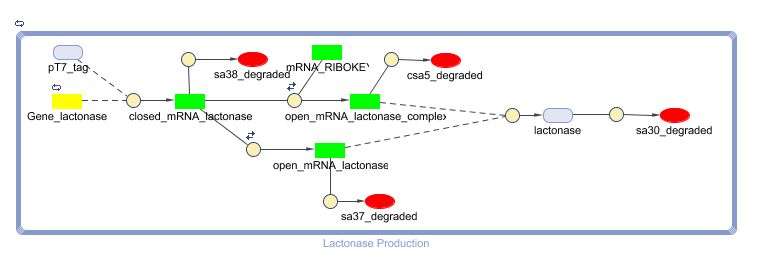
Constant Lactonase Production
Position in the system
The Constant Lactonase Production-system is directly linked to the Filter.
When the filter indicates that the input is zero (there is no desease), the system will (ideally) produce no lactonase. As soon as the output of the filter is one, the system starts producing lactonase and remains doing this untill the light goes off again. In this way all the HSL-molecules that are present will be 'removed' and the timer is reset.
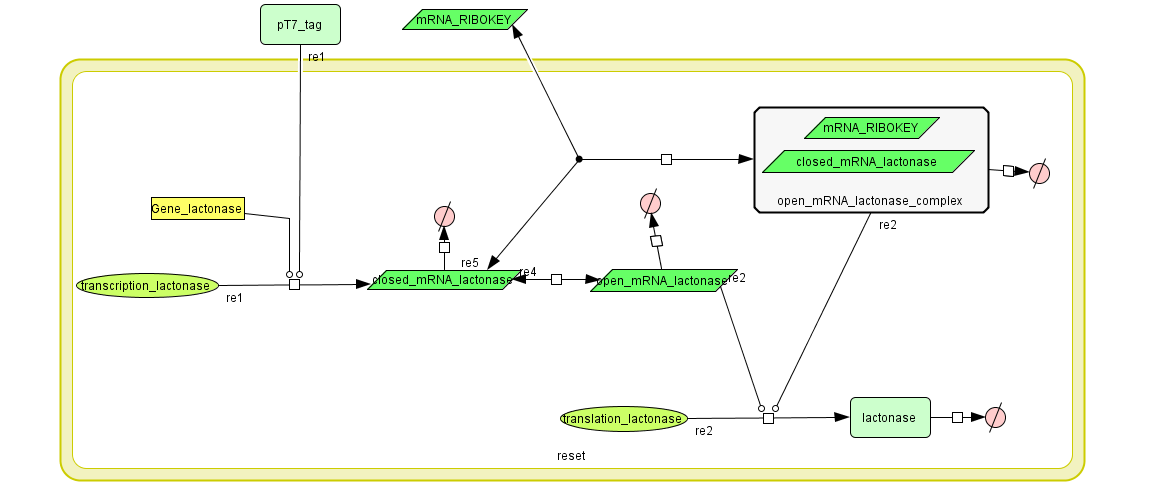
Describing the system
see also: Project:Reset
ODE's
Parameters
| Name | Value | Comments | Reference |
|---|---|---|---|
| Degradation rates | |||
| daiiA | dLVA = 2.814E-4 s-1 | LVA-tag reduces lifetime to 40 minutes | (3) |
| dclosed mRNA aiiA | 0.0046209812 s-1 | estimate: because this RNA isn't translated, it degrades faster | (2) |
| dopen mRNA aiiA | 0.0023104906 s-1 | (2) | |
| dmRNA aiiA complex | 0.0023104906 s-1 | (2) | |
| T7 Transcription | |||
| KT7 | 421 | dissociation constant, recalculated to remove units | (4) |
| kmax | 0.044 s-1 | maximal T7 transcription rate | (4) |
| Key-Lock constants | |||
| Keq 1 | 0,015 [M] | between closed and open Lactonase mRNA, modeled for competition, experimental | (1) |
| Keq 2 | 0.0212 [M] | between closed Lactonase mRNA and key unlocked mRNA complex, modeled for competition, experimental | (1) |
| kdis1 | 0.00416 s-1 | estimate: derived from experimental values | (1) |
| kcomplex1 | 0.00237 s-1 | estimate: derived from experimental values | (1) |
| kclosed | 500 s-1 | estimate: derived from experimental values | (1) |
| kopen | 7.5 s-1 | estimate: derived from experimental values | (1) |
| ktranslation | 0.167 s-1 | lock defined translation rate for Lactonase | (1) |
Models
CellDesigner (SBML file)
Matlab (SBML file)
Simulation
Remark: up to date with latest version?
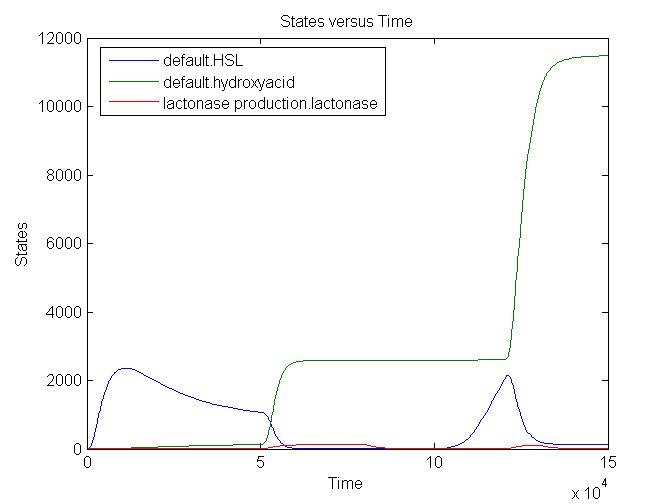
The number lactonase genes is held constant during the entire simulation. For the first 15.000 seconds the number of mRNA Ribokey is equal to 0.015 and the number of pT7 molecules to 0.4, then for 15000 s these numbers are set on 6 and 3 respectively (based on the results of the model of the filter) after which they are reduced back to 0.015 and 0.4.
We see that an increase in the number of mRNA Ribokey and pT7(due to an increase in light intensity) will lead to a much higher number of lactonase molecules.
References
| [1] | “Berkeley2006-RiboregulatorsMain - IGEM”; http://parts2.mit.edu/wiki/index.php/Berkeley2006-RiboregulatorsMain. |
| [2] | J.A. Bernstein et al., “Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays,” Proceedings of the National Academy of Sciences of the United States of America, vol. 99, Jul. 2002, pp. 9697–9702. |
| [3] | J.B. Andersen et al., “New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria,” Applied and Environmental Microbiology, vol. 64, Jun. 1998, pp. 2240–2246. |
| [4] | G.M. Skinner et al., “Promoter Binding, Initiation, and Elongation By Bacteriophage T7 RNA Polymerase: A SINGLE-MOLECULE VIEW OF THE TRANSCRIPTION CYCLE,” J. Biol. Chem., vol. 279, Jan. 2004, pp. 3239-3244. |
 "
"