Team:KULeuven/Project/CellDeath
From 2008.igem.org
(Difference between revisions)
m (→Action) |
|||
| Line 13: | Line 13: | ||
===Action=== | ===Action=== | ||
When the concentration of HSL is big enough (see the [https://2008.igem.org/Team:KULeuven/Model/CellDeath#CellDesigner_.28SBML_file.29 modeling page]), HSL will form a complex with the LuxR protein and thus activate the LuxpR promoter. This results in ''ccdB'' production and eventually in cell death ([http://www.ncbi.nlm.nih.gov/pubmed/10196173?dopt=Abstract reference]). | When the concentration of HSL is big enough (see the [https://2008.igem.org/Team:KULeuven/Model/CellDeath#CellDesigner_.28SBML_file.29 modeling page]), HSL will form a complex with the LuxR protein and thus activate the LuxpR promoter. This results in ''ccdB'' production and eventually in cell death ([http://www.ncbi.nlm.nih.gov/pubmed/10196173?dopt=Abstract reference]). | ||
| + | |||
| + | {{:Team:KULeuven/Tools/Components}} | ||
Revision as of 09:11, 1 August 2008
dock/undock dropdown
Contents |
Cell Death
BioBricks
Components
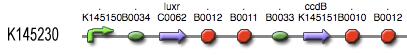
The luxR protein BBa_C0062 is placed under control of the constitutive promoter BBa_J23100. The ccdB suicide gene BBa_P1010, on its part, is placed under control of the LuxpR promoter BBa_R0062.
Action
When the concentration of HSL is big enough (see the modeling page), HSL will form a complex with the LuxR protein and thus activate the LuxpR promoter. This results in ccdB production and eventually in cell death (reference).
 "
"


 Input
Input Output
Output Filter
Filter InverTimer
InverTimer Reset
Reset Cell Death
Cell Death Memory
Memory