Team:UC Berkeley/GatewayPlasmid
From 2008.igem.org
(→Materials & Methods) |
(→General Procedure) |
||
| Line 35: | Line 35: | ||
== '''General Procedure''' == | == '''General Procedure''' == | ||
| + | |||
| + | Below is a flowchart of the general plasmid based Gateway procedure. | ||
== '''Results''' == | == '''Results''' == | ||
Revision as of 00:24, 29 October 2008
Contents |
Introduction
The current in vitro Gateway procedure for transferring one piece of DNA into another vector without restriction enzymes involves an expensive cocktail of purified plasmids, excisionase/integrase, ihfα and ihfβ. Both an assembly vector (containing the ccdB gene flanked by attR1 and attR2 sites for negative selection) and an entry vector (containing the part(s) desired to be transferred, flanked by attL1 and attL2 sites) are used as the substrates to which the reactants are added.
Inspired by such innovation, we seek to perform this task in vivo using E. coli to express the reactants necessary. Since E. coli naturally express ihfα/β in their genome, we first tried to integrate excise/integrase genes into the genomic DNA as well. However, this proved to be highly toxic to the cells (see Madhvi's Notebook). Therefore, our next approach was to introduce the reagents into the substrate plasmids. In one case, the the assembly vectors received the Gateway device of reagents excise/integrase preceded by a temperature sensitive promoter, while in the other case it was introduced into the entry vectors. In both cases, we also explored the effects of additionally introducing the ihfα and ihfβ genes behind the Gateway device.
After careful analysis of our data, we introduced our Lysis device in order to eliminate the mini-prepping procedures and to thus make the entire process cheaper and more efficient.
Gateway Device in Assembly Vector: Plasmid Details
Assembly Vectors:Two different variations of the pK112128 assembly vector were created: one with {promoter.xis.int!} and one with {promoter.xis.int!}{rbs.ihfα!}{rbs.ihfβ!} between the ccdB site and the attR2 site. These plasmids are named pK112245 and pK112246 respectively (see images below).
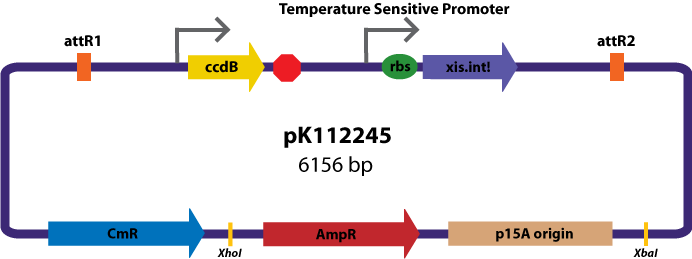
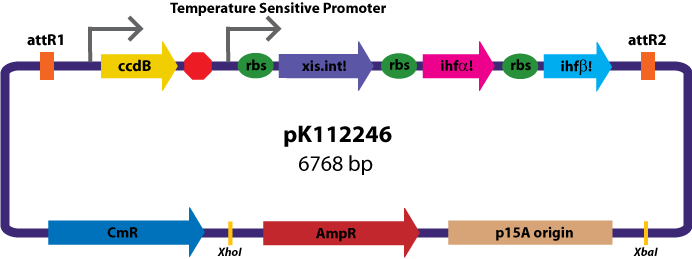
Entry Vector:The entry vector used in both cases was pBca1256 (see image below).
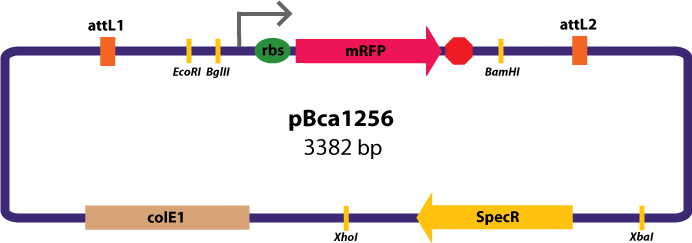
Gateway Device in Entry Vector: Plasmid Details
Assembly Vector:The assembly vector used for each of the following entry vector variations is pK112128 (see image below.
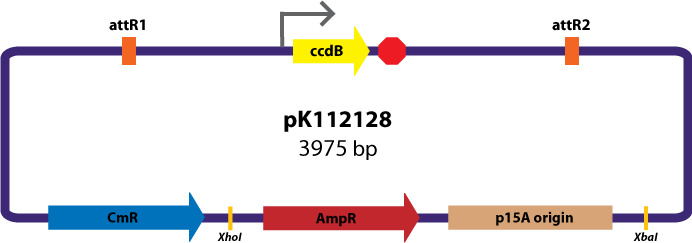
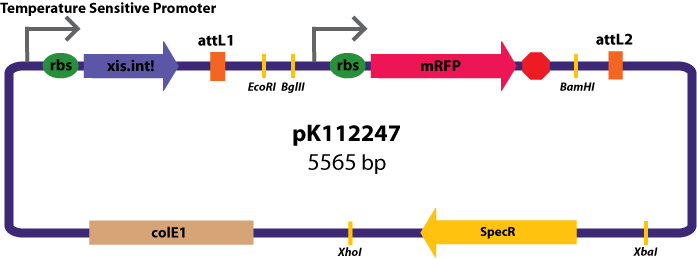
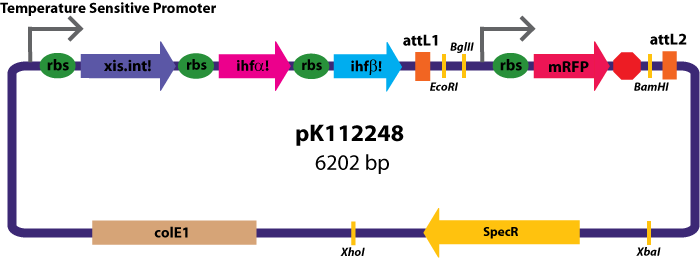
Entry Vectors: Two different variations of the pBca1256 entry vector were created: one with {promoter.xis.int!} and one with {promoter.xis.int!}{rbs.ihfα!}{rbs.ihfβ!} just before the attL1 site. These plasmids are named pK112247 and pK112248 respectively (see images below).
General Procedure
Below is a flowchart of the general plasmid based Gateway procedure.
Results
Materials & Methods
We first prepared competent TG1 cells (resistant to ccdB) with the given assembly vector. The assembly vector was transformed into competent TG1 cells and were grown in liquid LB media with Cam/Amp antibiotics (and 50mM Mg+2 if the lysis device is included) at 30°C overnight. The following day, 20 ul of the saturated culture was taken out, and was grown under the same conditions to mid-log. About 1 ml of culture was pelleted, the supernatant was discarded and the cells were resuspended in 10 ul KCM plus 90 ul TSS on ice.
Using these new competent cells, the given entry vector was tranformed into them. They were grown in liquid LB media with Cam/Amp/Spec antibiotics (and 50mM Mg+2 if the lysis device is included) at 37°C overnight to ensure the cells contain both plasmids and that the temperature sensitive promoter of the gateway device would turn on. A control in which the cells were instead kept at 30°C was conducted to compare our results with those in which the gateway device was theoretically off.
The following day, the plasmid from about 2 ml of saturated culture was purified. When there was no lysis device in the assembly vector, plasmid purification was achieved with the QIAprep Spin Miniprep Kit. In the case where the lysis device is integrated into the assembly plasmid, the culture was pelleted, the supernatant removed, resuspended in 1 ml PBS, pelleted, the supernatant removed, and finally resuspended again in 100 ul PBS.
The purified plasmids were next transformed into MG1061 cells (sensitive to ccdB). These were plated on LB agar plates with Cam/Amp antibiotics and grown overnight at 37°C. The next day, 16 colonies were picked from each plate, spotted on a Cam/Amp antibiotic plate, as well as Spec antibiotic plate to test for cotransformance of the entry vector with the desired product.
The above procedure were conducted in parallel with different combinations of assembly and entry vectors. The following combinations of assembly and entry vectors were done: pK112245 + pBca1256, pK112246 + pBca1256, pK112128 + pK112247, and pK112128 + pK112248. Each combination included a control in which the Gateway device was not turned on during the allotted time for the reaction to occur (i.e. they were grown at 30°C, when the temperature sensitive promoter is off). In addition to this, three trials of each were conducted. Finally, in order to see if gateway could be performed on parts of different sizes, five different parts in pBca1256 were chosen (Bca1117, Bca1133, Bca1270, Bca1252, and Bca1168) and tested with the assembly vector pK112245, as well as with pK112245. One colony from each was sequenced to ensure gateway had occurred by identification of attB1 and attB2 sites as well as the parts in the assembly vectors.
Other controls were also performed. Each assembly vector used was also transformed into into MG1061 cells which were put on Cam/Amp antibiotic LB agar plates to make sure ccdB was sufficient for negative selection. Only a very few number of white colonies grew for each, which was expected since it is known that ccdB is prone to mutation and weakening. The entry vectors were also each individually transformed into MC1061 cells and plated on Cam/Amp antibiotic plates. No colonies appeared, proving that the entry vector did not contain Cam/Amp resistance.
 "
"