Team:Hawaii/Notebook/2008-08-30
From 2008.igem.org
| Projects | Events | Resources | ||
|---|---|---|---|---|
| Sponsors | Experiments | Milestones | Protocols | |
| Notebook (t) | Meetings (t) |
Things we did today
Wetlab work
Construction of p+r and re-replacement of BB-pRL1383a MCS
- Grace
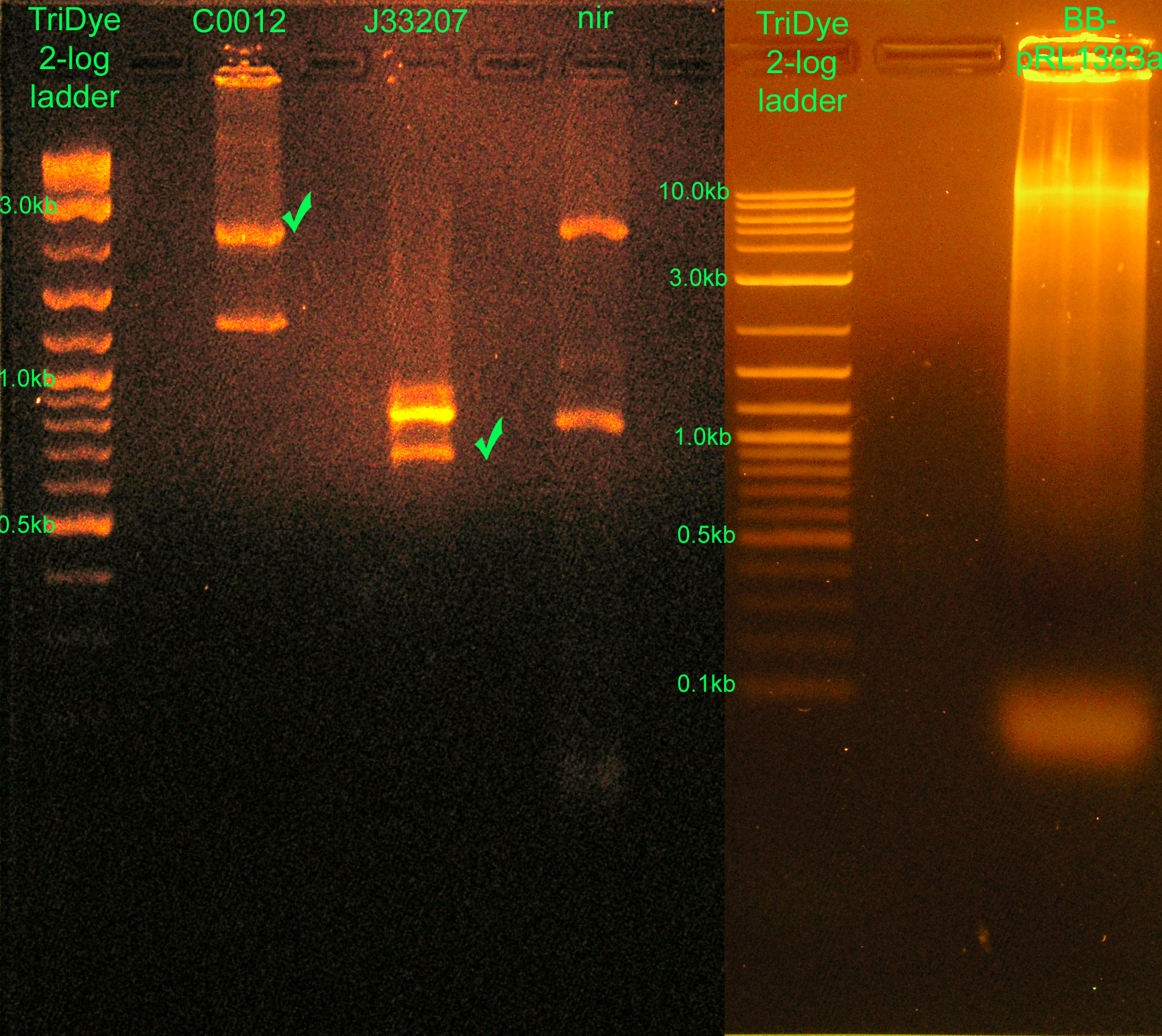
- Ran RE digests from last night on agarose gel
- C0012: correct vector band ~2.1kb
- J33207: band at ~700bp (only one enzyme cut -- XbaI did not cut; PstI cuts because C0012 was cut by PstI) and ~850bp (uncut PCR product)
- nir: ~800bp band (from GFP contaminant in plasmid prep, cut only once) and ~2kb band (??)
- BB-pRL1383a: ran WAY too much product. Will redo RE digest with 20% of plasmid used. Since J33207 was only cut once, it's assumed BB-pRL1383a was cut only once as well.
- Extracted C0012 vector band from gel
- Ligated:
- plac + rbs (B0034)
- PCR of J33207
- RE digested:
- p+r with EcoRI, XbaI, SpeI, PstI
- J33207 and BB-pRL1383a with EcoRI and PstI
- Treated C0012 vector (pSB1A2) with SAP
Construction of Broad-host-Range plasmid
- Margaret
File:PCR 8 30 08.jpg
PCR of rep, p1 omega.
- PCR:
- rep(50ul rxn), pRL1383a (template), rep(primers)
- P1 lytic(50ul rxn), pSB2K3(extracted from BB paper), P1(primers)
- (+) control: Base Vector + VF2&VR, (-) control: water + VF2&VR
- Culture: 5mL Terrific Broth + amp100
- plac+B0030 (1:6):3, 4, 7
- plac+B0030 :3
- plac+B0034 (1:6):1
- plac+B0034 (1:3):3
PCR
- Margaret
- P1 lytic (50ul rxn, pSB2K3(extracted from BB paper), P1(primers), 0.1ug/ml BSA
- omega (25ul rxn) The last gel had a ladder (so does this reaction, see gel from tomorrow).
Dry Lab
Added parts to registry
- Margaret
- plac+B0030:
- plac+B0034:
- oriT: K125320
- P1 lytic: K125330
- oriV:K125340
- rep proteins:K125800
- rep+oriV
Discussion
Quote of the Day
History is the only laboratory we have in which to test the consequences of thought. - Étienne Gilson
 "
"