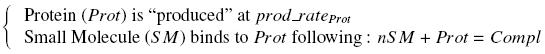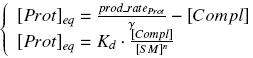Team:Paris/Modeling/Protocol Of Characterization
From 2008.igem.org
AnaJimenez (Talk | contribs) (→III-Promoter and Transcription Factors insertion) |
(→VI-B-Accessory plasmid construction) |
||
| (44 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
== I-Principles of the Experiments == | == I-Principles of the Experiments == | ||
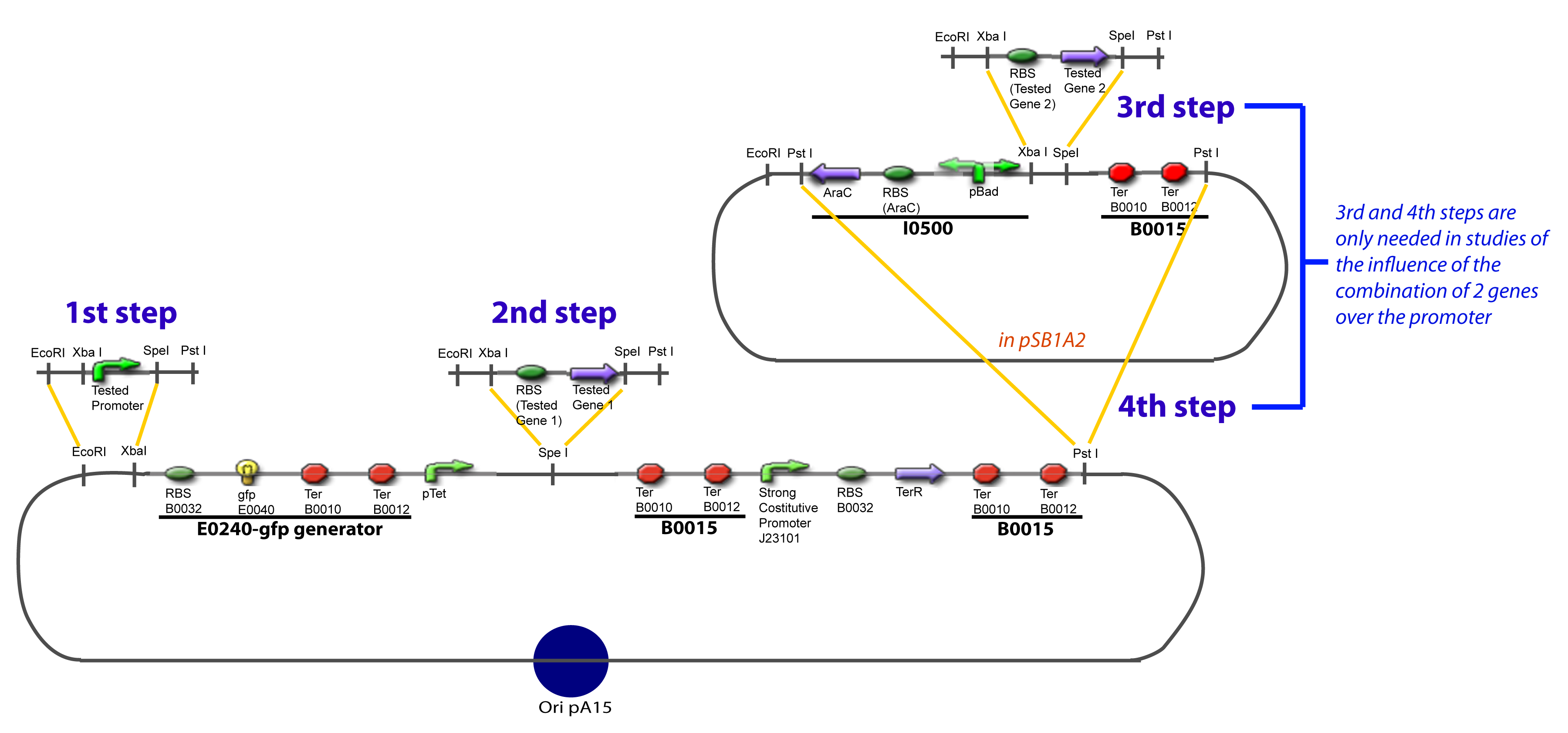
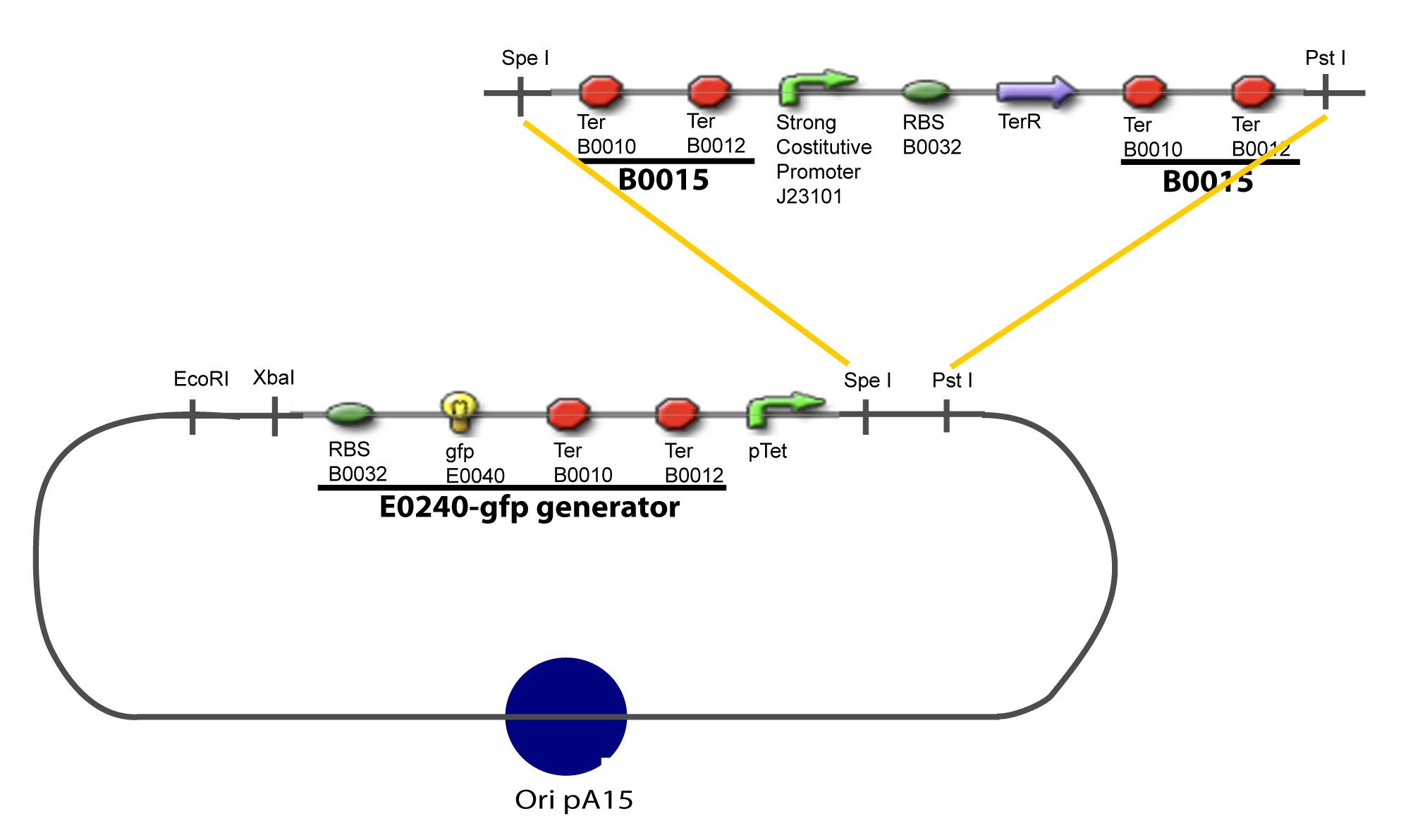
| - | To evaluate quantitativly the activity of a promoter in function of its transcription factors, we need data in which the different values of the activities are correlated with various ''known'' and ''controlled'' values of the transcription factors concentrations. Therefore, we designed a generic plasmid in which the transcription factors are put under the control of ''previously characterized inducible promoter'', and the studied promoter is put before a fluorescent reporter gene. In order to allow the study of the influence of two transcription factors over the tested pormoter, we chose to put the tested | + | To evaluate quantitativly the activity of a promoter in function of its transcription factors, '''we need data in which the different values of the activities are correlated with various ''known'' and ''controlled'' values of the transcription factors concentrations'''. <br><br> |
| + | Therefore, we designed a generic plasmid in which the transcription factors are put under the control of ''previously characterized inducible promoter'', and the studied promoter is put before a fluorescent reporter gene. In order to allow the study of the influence of two transcription factors over the tested pormoter, we chose to put the tested transcription factor (also called the "tested gene" in the following) under two different inducible systems . One is the ''pBAD-AraC'' system. The second one is an indirect system were the gene is after the Tet inducible promoter ''pTet''. <br><br> | ||
| + | The ''TetR'' gene would be expressed constitutively and at high rate thanks to a strong promoter (''J23101'') and its influence over the ''pTet'' promoter would be regulated by the concentration of the aTc molecule. That way, the production of the tested transcription factor can also be regulated, because the ''J23101'', and the ''pTet'' have been previously characterized. In a quite similar way, we characterize the ''pBad'' promoter.<br><br> | ||
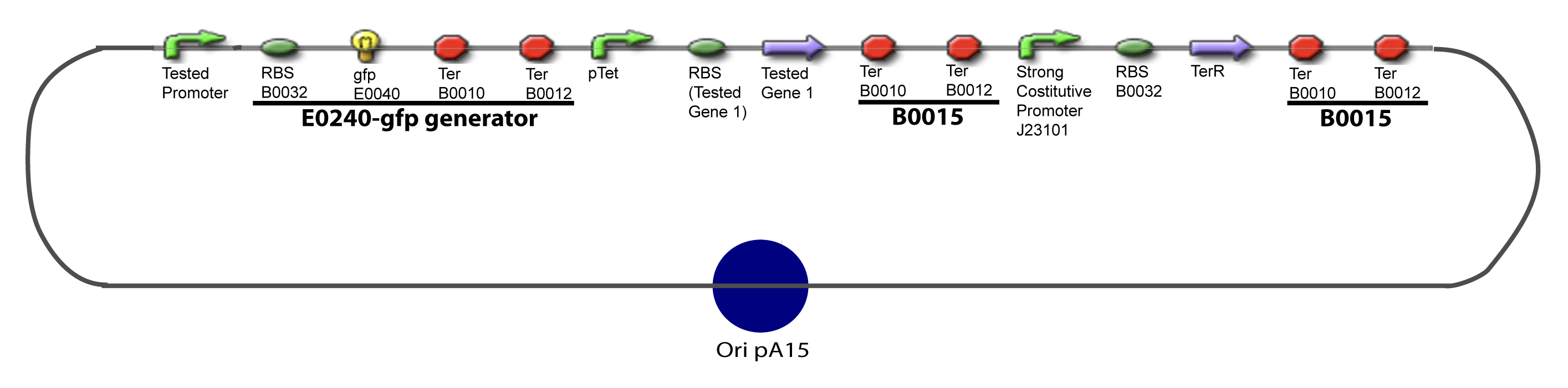
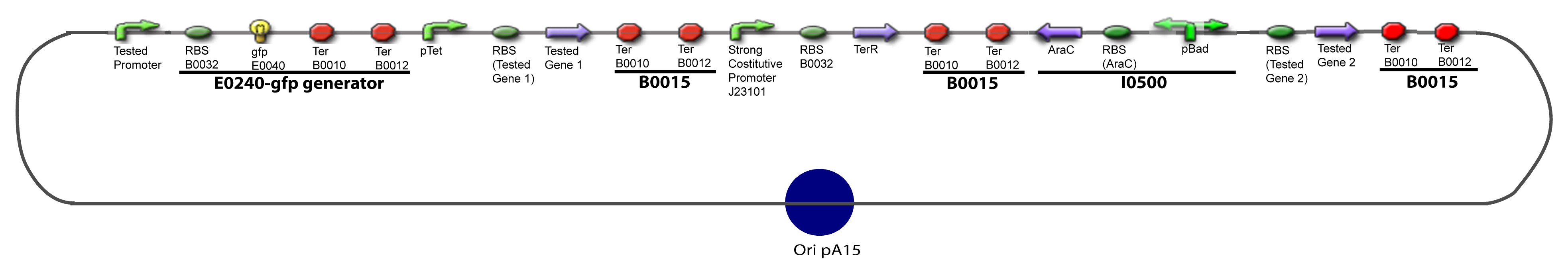
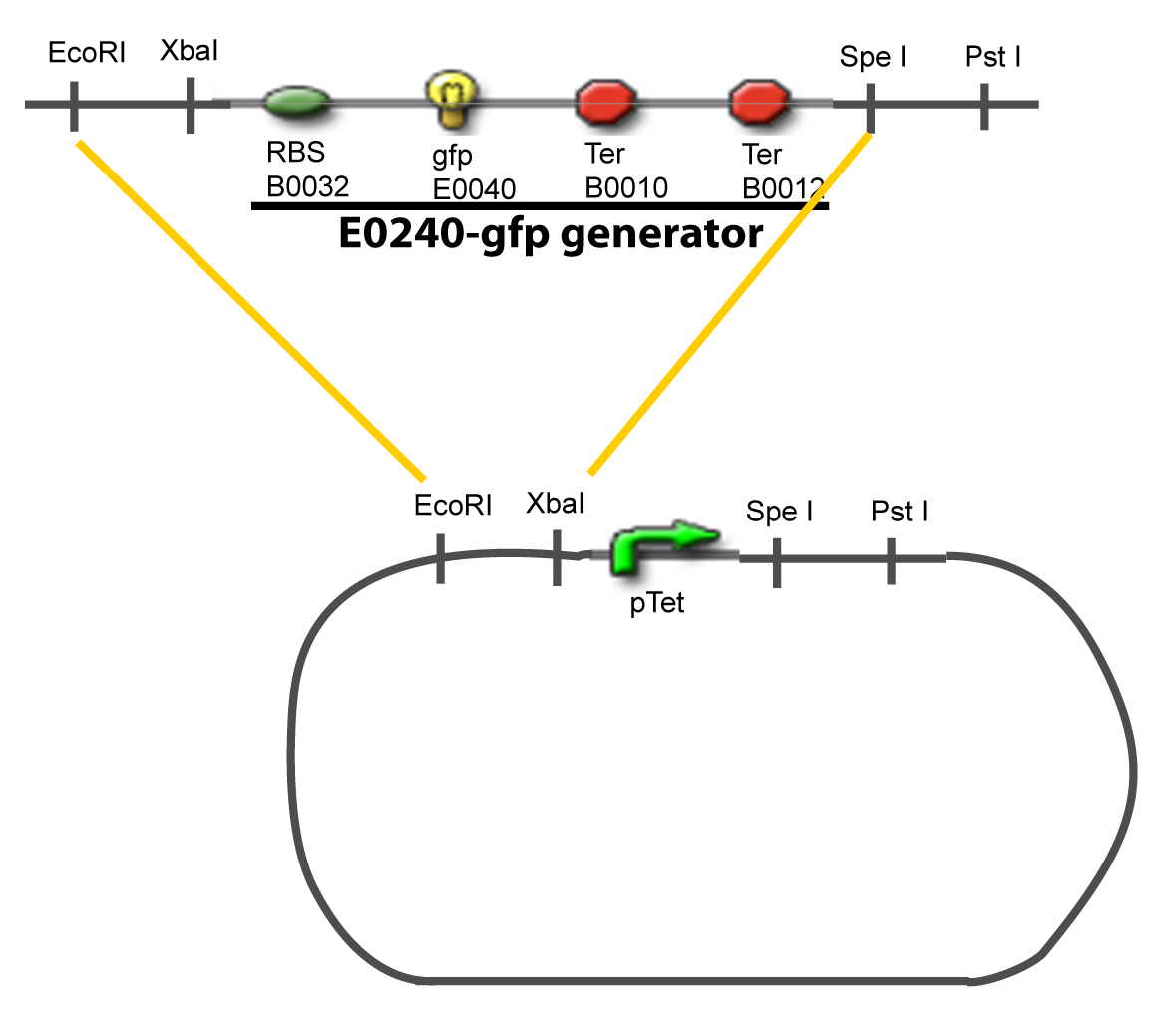
Here we show the design of two plasmids : one to test the influence of one gene and the other to test the influence of two genes over the tested promoter. | Here we show the design of two plasmids : one to test the influence of one gene and the other to test the influence of two genes over the tested promoter. | ||
| Line 13: | Line 15: | ||
[[Image:I-A-PCP.png|900px|The main design.]] | [[Image:I-A-PCP.png|900px|The main design.]] | ||
| - | |||
===II-B-For study with two Transcription Factor=== | ===II-B-For study with two Transcription Factor=== | ||
| Line 21: | Line 22: | ||
==III-Molecular design for Promoter Characterization Plasmid== | ==III-Molecular design for Promoter Characterization Plasmid== | ||
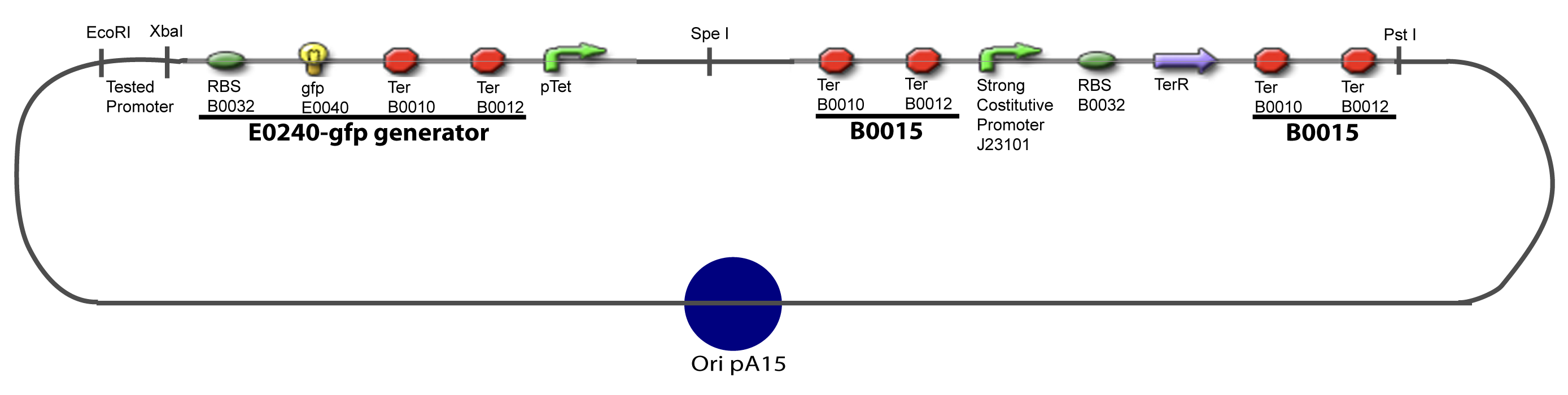
| - | Our aim is to make this plasmid useful not only for our project but for the whole iGEM comunity. This is why we decided to keep the Biobrick spirit as much as we could, making the plasmid compatible with the parts, so the teams using it | + | Our aim is to make this plasmid useful not only for our project but for the whole iGEM comunity. This is why we decided to keep the Biobrick spirit as much as we could, making the plasmid compatible with the parts, so the teams using it needs only the four traditional enzymes: EcoRI, XbaI, SpeI and PstI.We wanted also to make optional the introduction of a second tested transcription factor. The strategy is then based in two plasmids. <br><br> |
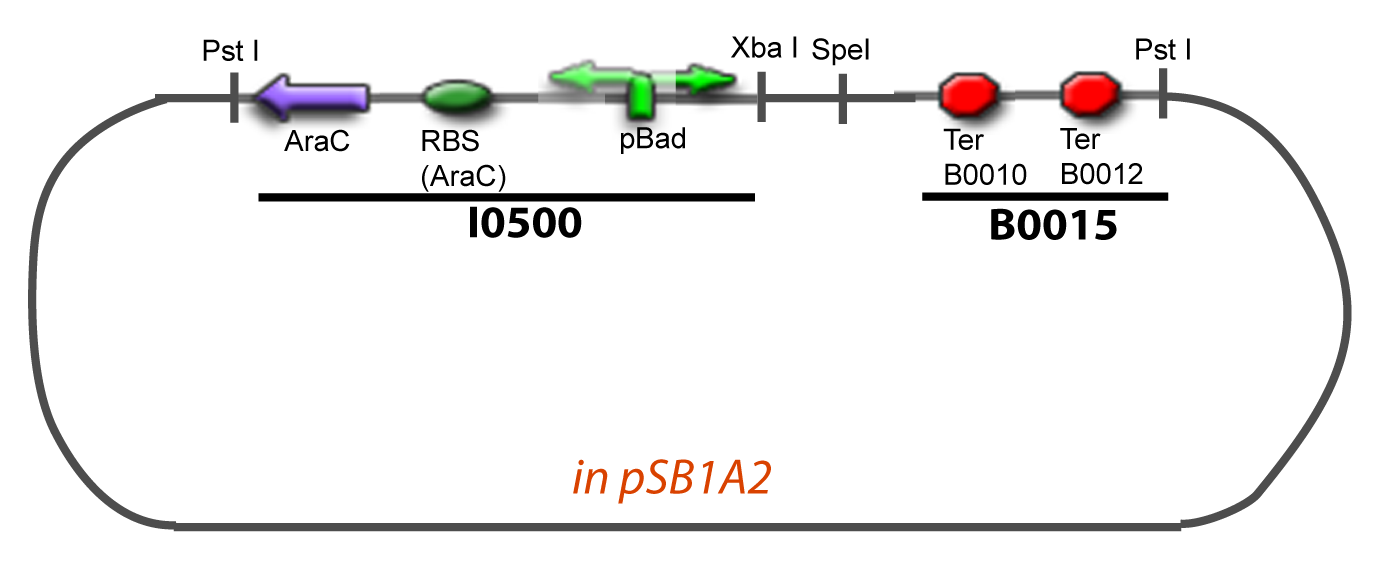
| + | The principal plasmid contains everything needed to test the effect of one gene over the tested promoter activity. The second plasmid called « Accessory plasmid » can be introduced easily in the Principal Plasmid and contains the necessary elements to add the expression of a second gene to the system. The resulting plasmids are presented below. | ||
| + | <div style="text-align: left"> | ||
| + | {{Paris/Toggle|'''III-A-Principal plasmid'''|Team:Paris/Modeling/More_Principal_Plasmid|900px}} | ||
| + | </div> | ||
| - | |||
| - | |||
| - | + | <div style="text-align: left"> | |
| - | + | {{Paris/Toggle|'''III-B-Accessory plasmid'''|Team:Paris/Modeling/More_Accessory_Plasmid|600px}} | |
| - | + | </div> | |
| - | + | ||
==IV-Promoter and Transcription Factors insertion== | ==IV-Promoter and Transcription Factors insertion== | ||
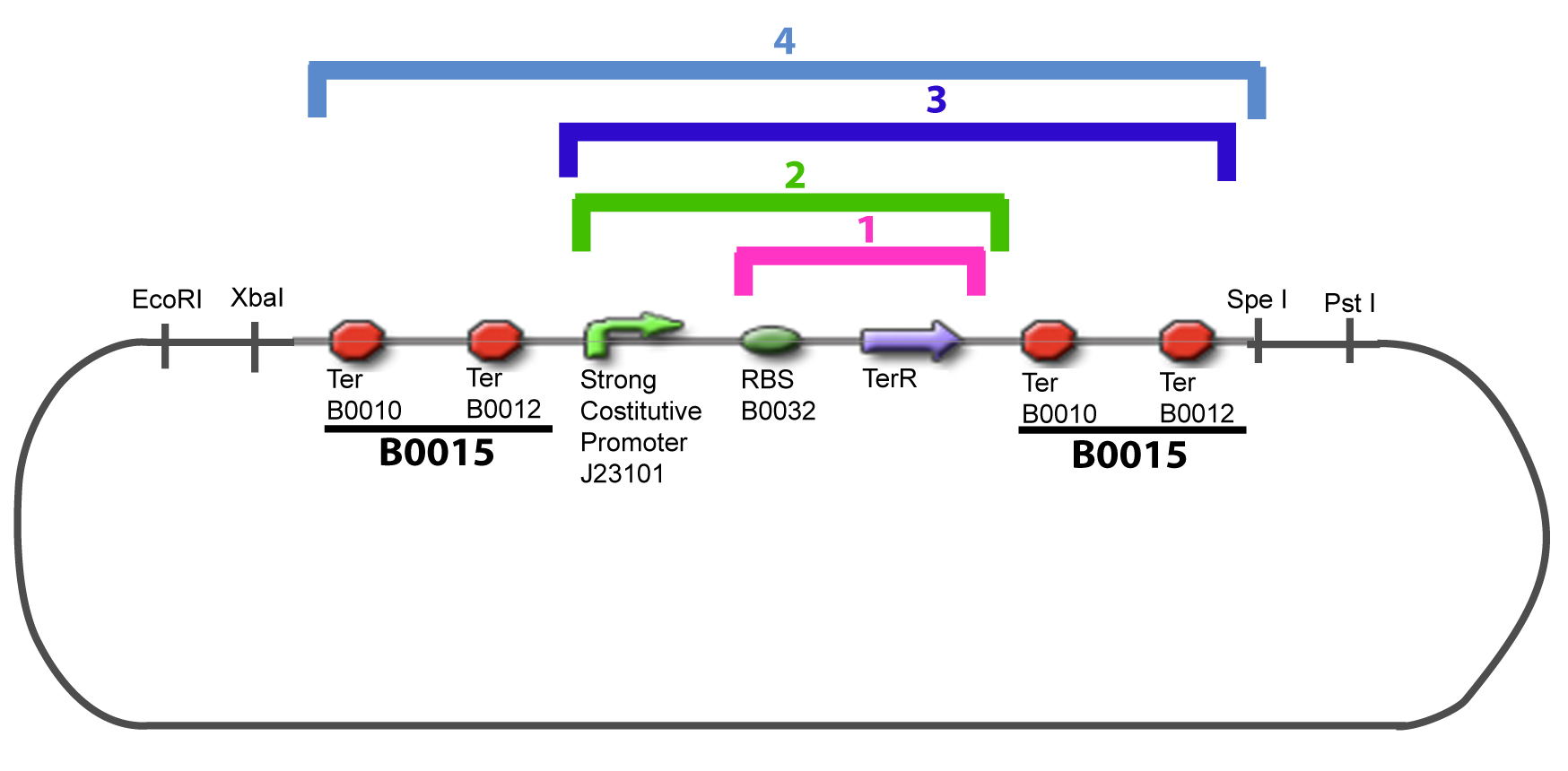
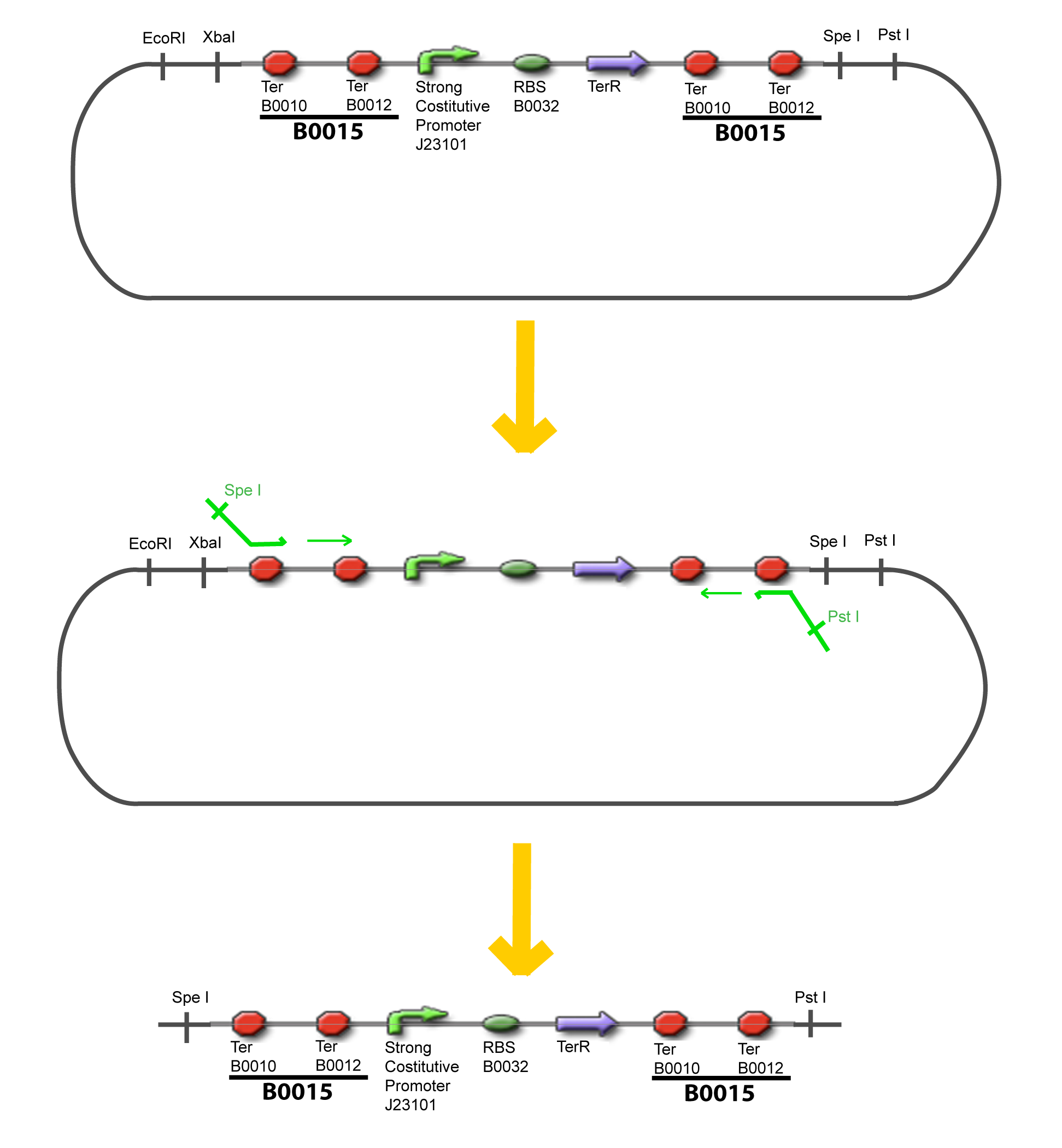
| - | The strategy to introduce the tested promoter and the tested | + | The strategy to introduce the tested promoter and the tested transcription factor(s) is very simple. The only difficulty is that the order of insertion that we describe has to be respected to avoid unwanted restriction enzyme cuts: |
| - | + | <div style="text-align: center"> | |
| - | + | {{Paris/Toggle|'''The Steps'''|Team:Paris/Modeling/More_The_Steps|700px}} | |
| - | + | </div> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | |||
| - | + | <div style="text-align: center"> | |
| - | + | {{Paris/Toggle|'''Illustration'''|Team:Paris/Modeling/More_Illustration_Steps|900px}} | |
| - | + | </div> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | ==V-Resulting plasmids== | ||
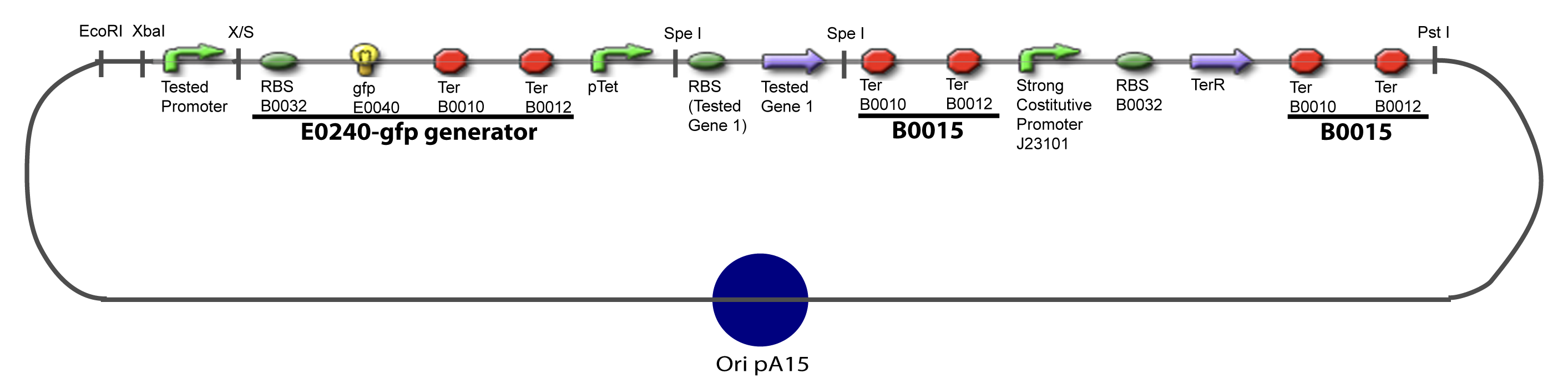
| - | + | ===V-A-For one transcription factor effect=== | |
| - | + | ||
| - | == | + | |
| - | + | ||
| - | + | ||
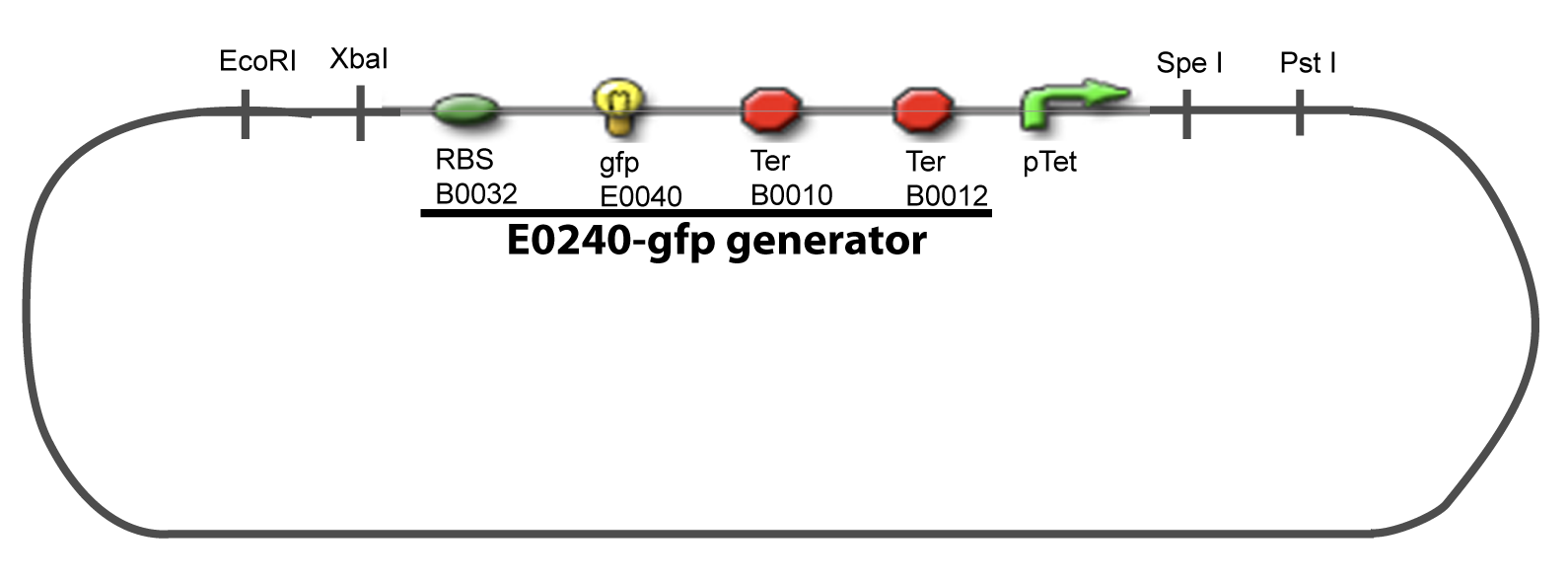
[[Image:IV-A-PCP.png|900px|The main design.]] | [[Image:IV-A-PCP.png|900px|The main design.]] | ||
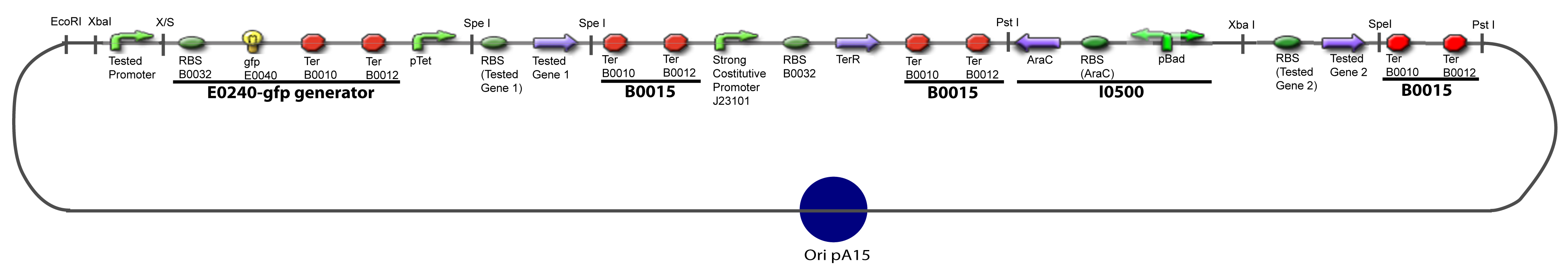
| - | === | + | ===V-B-For two transcription factors effect=== |
[[Image:IV-B-PCP.png|900px|The main design.]] | [[Image:IV-B-PCP.png|900px|The main design.]] | ||
| - | + | ==VI-Protocole for plasmid construction== | |
| - | == | + | |
We show here our plan to make these plasmids at the experimental level. | We show here our plan to make these plasmids at the experimental level. | ||
| - | === | + | ===VI-A-Principal Plasmid construction=== |
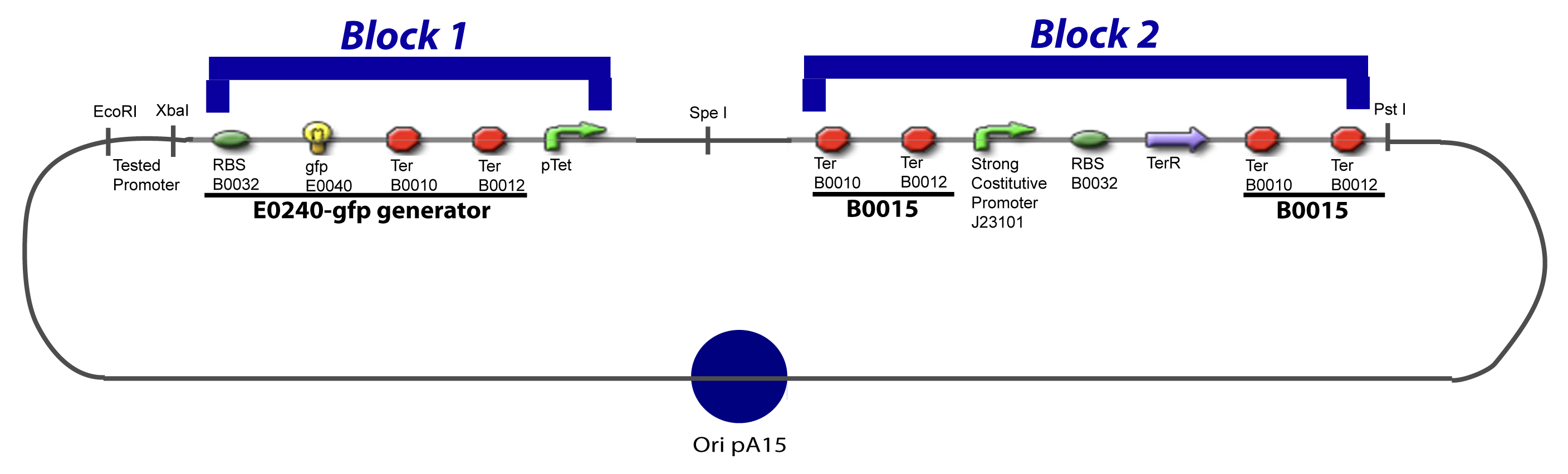
We devided the Principal Plasmid in two main blocks to go faster: | We devided the Principal Plasmid in two main blocks to go faster: | ||
| - | ==== | + | ====VI-A-1-Blocks==== |
[[Image:V-A-1-PCP.png|900px|The main design.]] | [[Image:V-A-1-PCP.png|900px|The main design.]] | ||
| - | = | + | <div style="text-align: left"> |
| + | {{Paris/Toggle|'''VI-A-2-Block 1 construction'''|Team:Paris/Modeling/More_Block_I|900px}} | ||
| + | </div> | ||
| - | + | <br> | |
| - | + | <div style="text-align: left"> | |
| + | {{Paris/Toggle|'''VI-A-3-Block 2 construction'''|Team:Paris/Modeling/More_Block_II|900px}} | ||
| + | </div> | ||
| - | + | <br> | |
| - | = | + | <div style="text-align: left"> |
| + | {{Paris/Toggle|'''VI-A-4-Two-block assembly'''|Team:Paris/Modeling/More_Blocks_Assembly|900px}} | ||
| + | </div> | ||
| - | + | <br><br> | |
| - | + | ===VI-B-Accessory plasmid construction=== | |
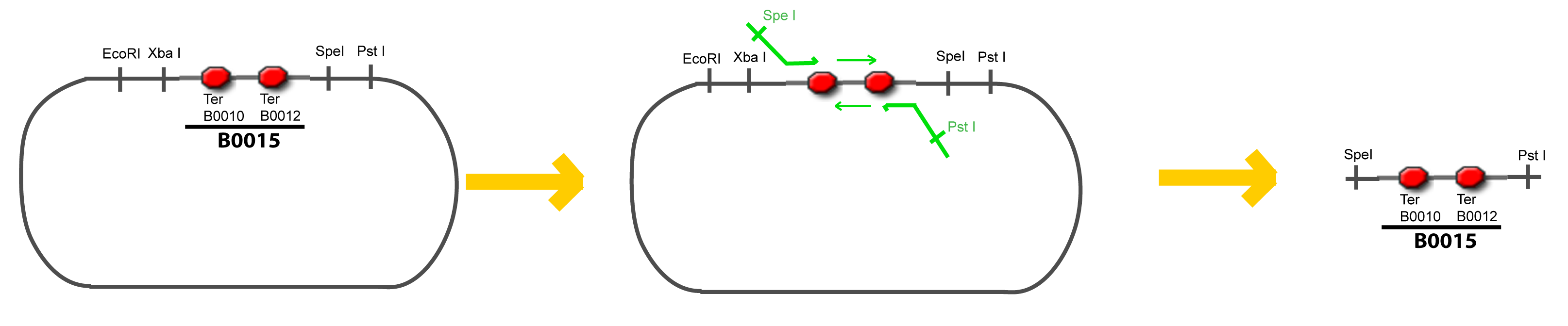
| + | Addition of restriction sites to B0015 by PCR: | ||
| - | + | <div style="text-align: center"> | |
| + | {{Paris/Toggle|Scheme|Team:Paris/Modeling/More_Scheme1|800px}} | ||
| + | </div> | ||
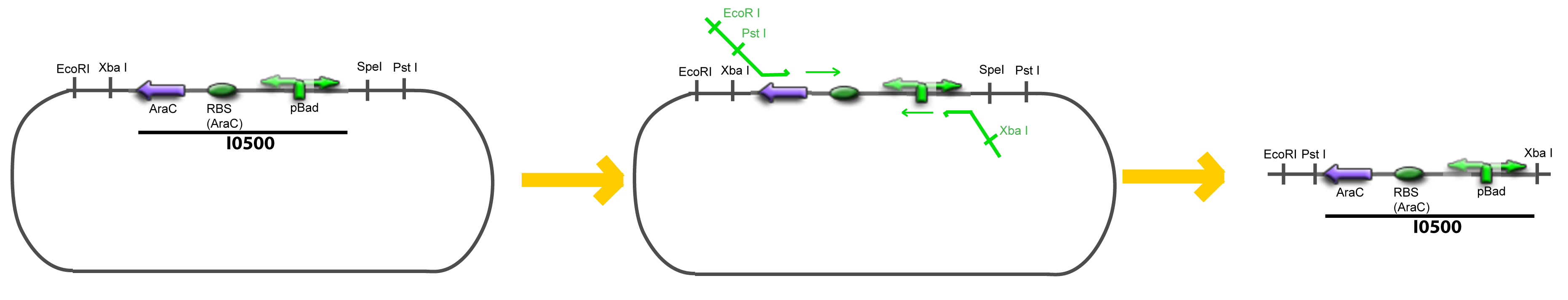
| - | + | Addition of restriction sites to ''pBAD-AraC'' by PCR: | |
| - | = | + | <div style="text-align: center"> |
| + | {{Paris/Toggle|Scheme|Team:Paris/Modeling/More_Scheme2|800px}} | ||
| + | </div> | ||
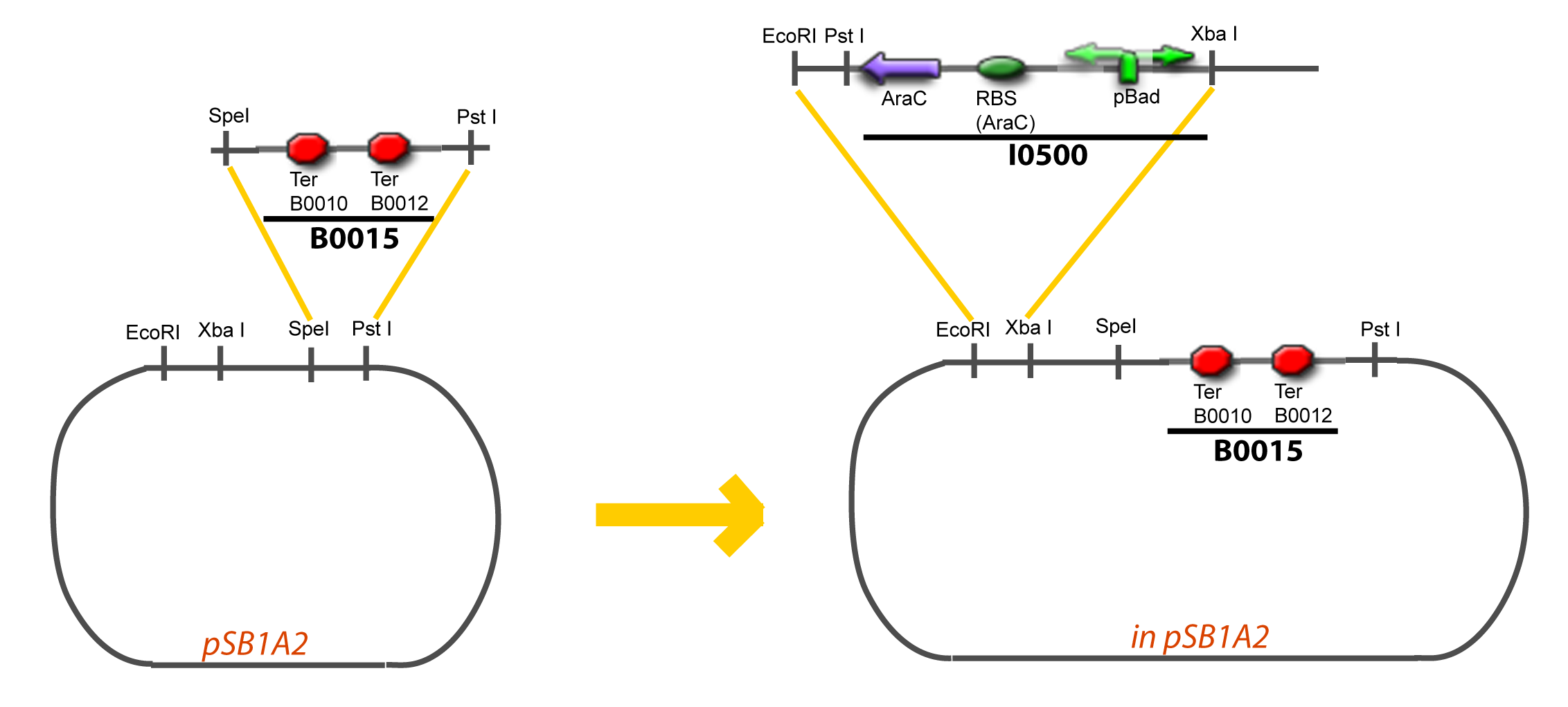
| + | Assembly: | ||
| - | + | <div style="text-align: center"> | |
| + | {{Paris/Toggle|Scheme|Team:Paris/Modeling/More_Scheme3|800px}} | ||
| + | </div> | ||
| - | == | + | == Specific Assumptions for the Experiments == |
| - | + | The previous experiment is used to find parameters regarding to our modelization. However, the experiments themselves must be described in the same way to interprete these parameters consistently. | |
| - | + | In this order, the two inductible promoters will be described exactly as the others (those in the system), with similar rules of complexation and transcription. | |
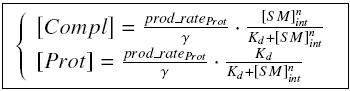
| - | + | Otherwise, as we control the quantity of transcription factor by the introduction of small molecules, we must model the diffusion/complexation/induction phenomena, which lead from the latter to the former. By putting the cell culture in a chemostat, we will be able to control precisely the concentration of the input small molecules. At ''quasi steady-state'', and assuming a ''simple, passive and quick diffusion'', we consider the '''inner concentration''', the '''outer medium concentration''', and the '''input concentration''' beeing all '''equal'''. | |
| - | + | <div style="text-align: center"> | |
| + | {{Paris/Toggle|Quick Mathematical Development|Team:Paris/Modeling/More_Math_Dvlp|700px}} | ||
| + | </div> | ||
| - | + | Thanks to these considerations, we can have access to these characterizations : | |
| + | |||
| + | <div style="text-align: center"> | ||
| + | {{Paris/Toggle|Prior characterizations|Team:Paris/Modeling/More_Algo_Prior|300px}} | ||
| + | </div> | ||
| - | + | and then | |
| - | = | + | <div style="text-align: center"> |
| + | {{Paris/Toggle|System characterizations|Team:Paris/Modeling/More_Algo_Finder|300px}} | ||
| + | </div> | ||
| - | + | <div style="text-align: center"> | |
| + | It is important to follow the previous chronological order for the successive characterizations, because some results are reused in the next ! | ||
| + | </div> | ||
<br> | <br> | ||
Latest revision as of 03:06, 30 October 2008
|
Protocol of Characterization
I-Principles of the ExperimentsTo evaluate quantitativly the activity of a promoter in function of its transcription factors, we need data in which the different values of the activities are correlated with various known and controlled values of the transcription factors concentrations. II-Plasmid for promoter characterizationII-A-For study with one Transcription FactorII-B-For study with two Transcription FactorIII-Molecular design for Promoter Characterization PlasmidOur aim is to make this plasmid useful not only for our project but for the whole iGEM comunity. This is why we decided to keep the Biobrick spirit as much as we could, making the plasmid compatible with the parts, so the teams using it needs only the four traditional enzymes: EcoRI, XbaI, SpeI and PstI.We wanted also to make optional the introduction of a second tested transcription factor. The strategy is then based in two plasmids.
IV-Promoter and Transcription Factors insertionThe strategy to introduce the tested promoter and the tested transcription factor(s) is very simple. The only difficulty is that the order of insertion that we describe has to be respected to avoid unwanted restriction enzyme cuts: ↓ The Steps ↑
V-Resulting plasmidsV-A-For one transcription factor effect
V-B-For two transcription factors effectVI-Protocole for plasmid constructionWe show here our plan to make these plasmids at the experimental level. VI-A-Principal Plasmid constructionWe devided the Principal Plasmid in two main blocks to go faster: VI-A-1-Blocks
VI-B-Accessory plasmid constructionAddition of restriction sites to B0015 by PCR: Addition of restriction sites to pBAD-AraC by PCR: Assembly: Specific Assumptions for the ExperimentsThe previous experiment is used to find parameters regarding to our modelization. However, the experiments themselves must be described in the same way to interprete these parameters consistently. In this order, the two inductible promoters will be described exactly as the others (those in the system), with similar rules of complexation and transcription. Otherwise, as we control the quantity of transcription factor by the introduction of small molecules, we must model the diffusion/complexation/induction phenomena, which lead from the latter to the former. By putting the cell culture in a chemostat, we will be able to control precisely the concentration of the input small molecules. At quasi steady-state, and assuming a simple, passive and quick diffusion, we consider the inner concentration, the outer medium concentration, and the input concentration beeing all equal. ↓ Quick Mathematical Development ↑
Thanks to these considerations, we can have access to these characterizations : ↓ Prior characterizations ↑
and then ↓ System characterizations ↑
It is important to follow the previous chronological order for the successive characterizations, because some results are reused in the next !
|
 "
"