Imperial College/29 August 2008
From 2008.igem.org
(Difference between revisions)
| Line 41: | Line 41: | ||
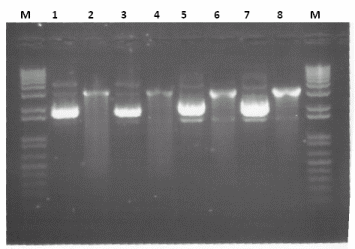
*'''Genomic''' - From the results the optimum conditions is clearly reaction 5 (10 cycles at 55<sup>o</sup>C, 20 cycles at 62<sup>o</sup>C, Primers = Fw1/R1) | *'''Genomic''' - From the results the optimum conditions is clearly reaction 5 (10 cycles at 55<sup>o</sup>C, 20 cycles at 62<sup>o</sup>C, Primers = Fw1/R1) | ||
*'''Single Colony PCR''' - From the results clearly it is sufficient to use only 1ul of sample. In addition only the combination of Fw1 and Rv1 is working at either ''10 cycles at 52<sup>o</sup>C, 20 cycles at 64<sup>o</sup>C'' or ''10 cycles at 50<sup>o</sup>C, 20 cycles at 64<sup>o</sup>C'' | *'''Single Colony PCR''' - From the results clearly it is sufficient to use only 1ul of sample. In addition only the combination of Fw1 and Rv1 is working at either ''10 cycles at 52<sup>o</sup>C, 20 cycles at 64<sup>o</sup>C'' or ''10 cycles at 50<sup>o</sup>C, 20 cycles at 64<sup>o</sup>C'' | ||
| + | <br> | ||
| + | {{Imperial/EndPage|Notebook|Notebook}} | ||
Latest revision as of 20:40, 28 October 2008
29 August 2008WetLabMidipreps
PCRIn a pursuit to verify the various PCR protocols and PCR primers we carried out the following reactions:
For each of these reactions a gradient of temperatures was used during the 10 cycles in the annealing stage of PCR and also the following 20 cycles, see our protocol for more information in summary the following reactions were carried out:
Results
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"