Team:TUDelft/Temperature design2
From 2008.igem.org
Bavandenberg (Talk | contribs) (→) |
Bavandenberg (Talk | contribs) (→Changing the temperature threshold of the RNA thermometer) |
||
| Line 17: | Line 17: | ||
But things are not that easy. At first, adding mutations in order to enforce or weaken the temperature sensitive region can also cause the RNA to fold into a completely different structure. This way it can loose its function as an RNA thermometer. Secondly, there are some constraints to the possible mutations; the start codon and the Shine Dalgarno sequence must off course remain unaltered and in case of a standard biobrick the scar must also be part of the temperature sensitive region (see the [https://2008.igem.org/Team:TUDelft/Temperature_design#Scar_problem scar problem]). Third, although it seems that only a thermosensitive hairpin is needed to have a functioning RNA thermometer (as described in the [https://2008.igem.org/Team:TUDelft/Temperature_analysis#What_is_needed_to_have_a_functioning_RNA_thermometer analysis section]) there could very well be more factors that are of influence on the functioning and temperature threshold of the RNA thermometer that are still unknown. | But things are not that easy. At first, adding mutations in order to enforce or weaken the temperature sensitive region can also cause the RNA to fold into a completely different structure. This way it can loose its function as an RNA thermometer. Secondly, there are some constraints to the possible mutations; the start codon and the Shine Dalgarno sequence must off course remain unaltered and in case of a standard biobrick the scar must also be part of the temperature sensitive region (see the [https://2008.igem.org/Team:TUDelft/Temperature_design#Scar_problem scar problem]). Third, although it seems that only a thermosensitive hairpin is needed to have a functioning RNA thermometer (as described in the [https://2008.igem.org/Team:TUDelft/Temperature_analysis#What_is_needed_to_have_a_functioning_RNA_thermometer analysis section]) there could very well be more factors that are of influence on the functioning and temperature threshold of the RNA thermometer that are still unknown. | ||
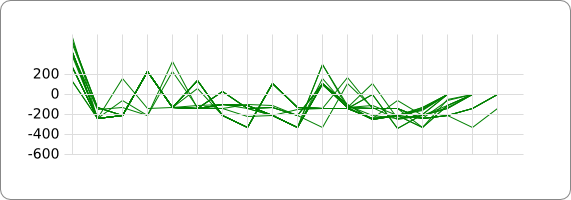
| - | == | + | [[Image:Tudelft summed unaligned noloop.png]] |
| + | |||
| + | |||
| + | |||
| + | == blaat == | ||
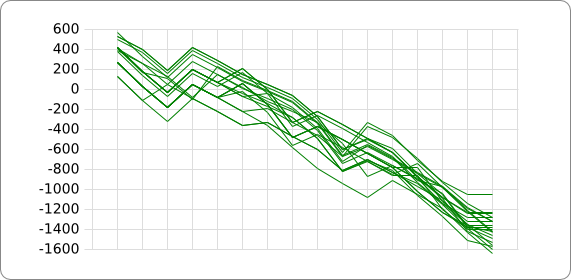
[[Image:Tudelft_notsummed_aligned.png | thumb]] | [[Image:Tudelft_notsummed_aligned.png | thumb]] | ||
Revision as of 08:09, 2 September 2008
>> Work in progress
BBa:K115016, BBa:K115017, BBa:K115018, BBa:K115019
Changing the temperature threshold of the RNA thermometer
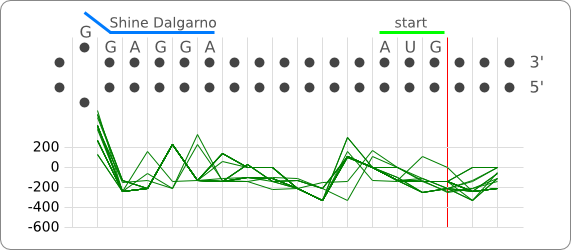
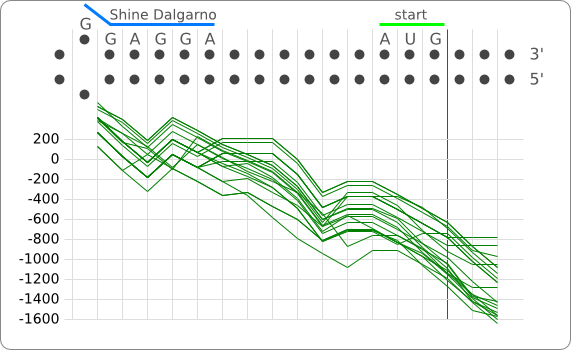
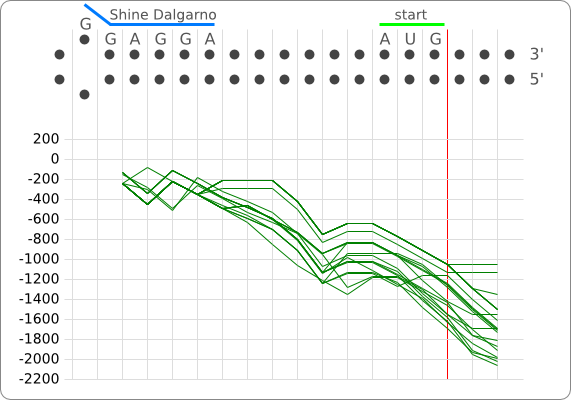
The principle of the RNA thermometer is based on base pairing between the nucleotides in the Shine Dalgarno region. At a certain temperature the RNA is folded into a structure in which base pairing nucleotides make the Shine Dalgarno region unreachable for the ribosome. With a rise in temperature, the binding forces between the base pairing nucleotides decrease and above a certain threshold temperature the base pairing forces are to weak to hold the base pairing nucleotides together, causing the RNA to (partially) unfold. The binding between the base pairing nucleotides let loose causing the RNA to unfold, exposing the Shine-Dalgarno region, enabling the ribosome to initiate translation.
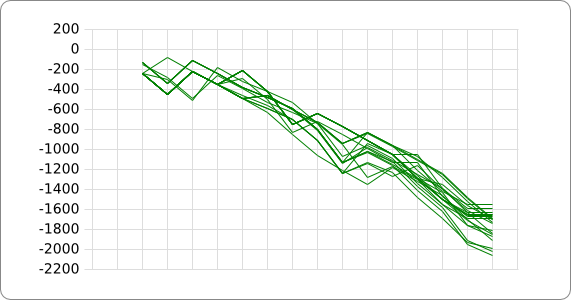
When only looking at this principle, an RNA thermometer with a different temperature threshold can be designed by simply increasing or decreasing the binding forces of the base-pairing nucleotides in the Shine-Dalgarno region, shifting the temperature threshold to a higher or lower temperature respectively (fig). Increasing the binding forces could be achieved by incorporating base pairing C and G nucleotides, which bind relatively strong forming a stable helix. Decreasing the binding forces could be done by introducing less strong binding nucleotide base-pairs, such as A-U and G-U base pairs, forming a less stable helix.
But things are not that easy. At first, adding mutations in order to enforce or weaken the temperature sensitive region can also cause the RNA to fold into a completely different structure. This way it can loose its function as an RNA thermometer. Secondly, there are some constraints to the possible mutations; the start codon and the Shine Dalgarno sequence must off course remain unaltered and in case of a standard biobrick the scar must also be part of the temperature sensitive region (see the scar problem). Third, although it seems that only a thermosensitive hairpin is needed to have a functioning RNA thermometer (as described in the analysis section) there could very well be more factors that are of influence on the functioning and temperature threshold of the RNA thermometer that are still unknown.
blaat
Two facts that enforce this assumption... First, aligning the sequences of the different types of RNA thermometers produces conserved regions around the SD region. Second, it is shown by De Smit and Van Duin (reference) that there is a strong correlation between translation efficiency and the stability of the helix containing the Shine-Dalgarno region and the initiation codon.
But will this region on itself be enough to have a working RNA thermometer? Looking at the consensus structure of the ROSE and PrfA RNA thermometer, it can be seen that there are other regions within the structure that are highly conserved. This indicates that these regions might also be of importance to the functioning of the RNA thermometer. It is still unknown if the stem-loops not containing the SD are of any help with the opening of the SD region (reference).
Pseudoknot analysis.
figures...(consensus structures, highly conserved SD regions)
De Smit and Van Duin [1] show a correlation between the level of expression and the free energy of the secondary structure in the Shine Dalgarno region.
References
- ^ De Smit M H, Van Duijn J (1990). "Secondary structure of the ribosome binding site determines translation efficiency: A quantitative analysis". PNAS, 1990-10, vol.87, no.19, 7668-7672. PMID:2217199
- ^ Hoe N P, Goguen J D (1993). "Temperature sensing in Yersinia pestis: Translation of the LcrF activator protein is thermally regulated". J Bacteriol, 1993 December, 175(24), 7901-7909. PMID:7504666
- ^ Chowdhurry S, Maris C, Allain F H T, Narberhaus F (2006). "Molecular basis for temperature sensing by an RNA thermometer". The EMBO Journal, 2006, 25, 2487–2497. PMID:16710302
- ^ Nocker A, Hausherr T, Balsiger S, Krstulovic N, Hennecke H, Narberhaus F (2001). "A mRNA-based thermosensor controls expression of rhizobial heat shock genes". Nucleic Acids Research, 2001 December 1, 29(23):4800-4807. PMID:11726689
- ^ Balsiger S, Ragaz C, Baron C, Narberhaus F. "Replicon-specific regulation of small heat shock genes in Agrobacterium tumefaciens". Journal of Bacteriology, October 2004, p. 6824-6829, Vol.186, No.20. PMID:15466035
- ^ Waldminghaus T, Heidrich N, Branti S, Narberhaus F (2007). "FourU: a novel type of RNA thermometer in Salmonella". Molecular Microbiology, Volume 65, Issue 2, 413-424. PMID:17630972
- ^ Johansson J, Mandin P, Renzoni A, Chiaruttinni C, Springer M, Cossart P. "An RNA thermosensor controls expression of virulance genes in Listeria monocytogenes". Cell , Volume 110 , Issue 5 , 551-561. PMID:12230973
 "
"