Imperial College/3 September 2008
From 2008.igem.org
(Difference between revisions)
m (→Transformation of B.subtilis) |
|||
| (11 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
{| cellpadding="10" border="0" | {| cellpadding="10" border="0" | ||
|- valign="top" | |- valign="top" | ||
| - | |||
|{{#calendar: title=Imperial_College |year=2008 | month=08}} | |{{#calendar: title=Imperial_College |year=2008 | month=08}} | ||
|{{#calendar: title=Imperial_College |year=2008 | month=09}} | |{{#calendar: title=Imperial_College |year=2008 | month=09}} | ||
| - | | rowspan="2" bgcolor=# | + | |{{#calendar: title=Imperial_College |year=2008 | month=10}} |
| + | | rowspan="2" bgcolor=#ffffff width="100%" | | ||
|} | |} | ||
| + | =3 September 2008= | ||
==Wetlab== | ==Wetlab== | ||
| + | |||
| + | ===Cloning=== | ||
| + | |||
| + | *Results of PCR reactions run on the 2nd analysed on a gel (results below - top gel) | ||
| + | *To check that PCR with Pfu was possible, a PCR reaction with the strongest vector PCR with taq was performed with Pfu under standard Pfu conditions[[http://openwetware.org/wiki/IGEM:IMPERIAL/2008/Prototype/Wetlab/PCR]] but a longer annealing time (1 minute); the primers used were AmyE 5' forward and reverse primers at 52°C for the first 10 cycles then 60°C for 20 cycles. Aad9 PCR was again tested according to the standard protocol and agin at 52°C for the first 10 cycles then 58°C or 56°C for 20 cycles (results below - lower gel) | ||
| + | *GFP-Terminator biobrick and RFP-Terminator biobrick cut again with ''XbaI'' and ''SpeI'' to produce biobrick vectors for our PCR clones | ||
| + | *GFP-Terminator biobrick and RFP-Terminator biobrick cut again with ''EcoRI'' and ''SpeI'' to produce biobrick vectors for our GeneArt produced clones | ||
| + | *Biobrick digest run on a gel for gel purification of vector | ||
| + | *GeneArt constructs in vectors midiprepped from Xl1-Blue cells | ||
| + | |||
===Transformation of ''B.subtilis''=== | ===Transformation of ''B.subtilis''=== | ||
*Previously we have tried to test efficient integration into the ''B.subtilis'' genome. However, the primers we were using have so far failed to yield either a positive or negative result when a signle colony PCR was carried out on transformed ''B.subtilis''. In order to check firstly if the primers are working and secondly if the single colony PCR protocol is working we carried out a series of testing. First to test the verification primers we are going to carry out a PCR on purified genome, that we have previously used successfully in a PCR reaction. We set up the following conditions: | *Previously we have tried to test efficient integration into the ''B.subtilis'' genome. However, the primers we were using have so far failed to yield either a positive or negative result when a signle colony PCR was carried out on transformed ''B.subtilis''. In order to check firstly if the primers are working and secondly if the single colony PCR protocol is working we carried out a series of testing. First to test the verification primers we are going to carry out a PCR on purified genome, that we have previously used successfully in a PCR reaction. We set up the following conditions: | ||
| Line 30: | Line 41: | ||
===Results=== | ===Results=== | ||
| - | [[Image: | + | [[Image:PCR 2-8.PNG|thumb|600px|center|A 1% Agarose gel showing the results of various PCR reactions]] |
| + | Upper Lanes: | ||
| + | *Lane 1- Pfu PCR reaction, first 10 cycles at 52<sup>o</sup>C and then 20 cycles at 60<sup>o</sup>C using AmyE 5' integration sequence primers, (effectively a positive control) | ||
| + | *Lane 2- Pfu PCR reaction, first 10 cycles at 50<sup>o</sup>C and then 20 cycles at 65<sup>o</sup>C using AmyE 3' integration sequence primers, | ||
| + | *Lane 3- Pfu PCR reaction, first 10 cycles at 48<sup>o</sup>C and then 20 cycles at 65<sup>o</sup>C using AmyE 3' integration sequence primers, | ||
| + | *Lane 4- Pfu PCR reaction, first 10 cycles at 56<sup>o</sup>C and then 20 cycles at 65<sup>o</sup>C using LacI gene primers, | ||
| + | *Lane 5- Taq PCR reaction, first 10 cycles at 56<sup>o</sup>C and then 20 cycles at 65<sup>o</sup>C using LacI gene primers but no template DNA (negative control) | ||
| + | <br> | ||
| + | *Lane 6- Taq PCR reaction, first 10 cycles at 54<sup>o</sup>C and then 20 cycles at 60<sup>o</sup>C using Aad9 primers, | ||
| + | *Lane 7- Taq PCR reaction, first 10 cycles at 52<sup>o</sup>C and then 20 cycles at 60<sup>o</sup>C using Aad9 primers, | ||
| + | *Lane 8- Taq PCR reaction, first 10 cycles at 50<sup>o</sup>C and then 20 cycles at 60<sup>o</sup>C using Aad9 primers, | ||
| + | *Lane 9 - Taq PCR reaction, first 10 cycles at 50<sup>o</sup>C and then 20 cycles at 60<sup>o</sup>C using Aad9 primers but no template DNA | ||
| + | *As can be observed all the PCRs failed, including the positive control. The most likely explanation is that the reactions were left to long before PCR started (in the case of Pfu) or that a vital reaction component was not correctly added to the reactions. A trial of Pfu will need to be carried out to determine if the enzyme itself may be a problem | ||
| + | |||
| + | Lower Lanes: | ||
| + | *Lane 1 - Annealing step 60<sup>o</sup>C using verification primers, | ||
| + | *Lane 2- Annealing step 58<sup>o</sup>C using verification primers, | ||
| + | *Lane 3- Annealing step 56<sup>o</sup>C using verification primers, | ||
| + | *Lane 4- Annealing step 56<sup>o</sup>C using verification primers, no DNA negative control, | ||
| + | <br> | ||
| + | *Lane 5 - Annealing step 52<sup>o</sup>C using AmyE fw1 and rev2 primers, | ||
| + | *Lane 6- Annealing step 50<sup>o</sup>C using AmyE fw1 and rev2 primers, | ||
| + | *Lane 7- Annealing step 48<sup>o</sup>C using AmyE fw1 and rev2 primers, | ||
| + | *Lane 8- Annealing step 61<sup>o</sup>C using AmyE fw1 and rev1, positive control, | ||
| + | *Lane 9 - Annealing step 48<sup>o</sup>C using AmyE fw1 and rev2, no DNA negative control, | ||
| + | <br> | ||
| + | <br> | ||
| + | *As we can see only for the positive control we see a band, this means that the verfication primers and the fw1 and rev 2 combinations are not working. We will double check the primer sequences and carry on trying to optimise the protocol. | ||
| + | |||
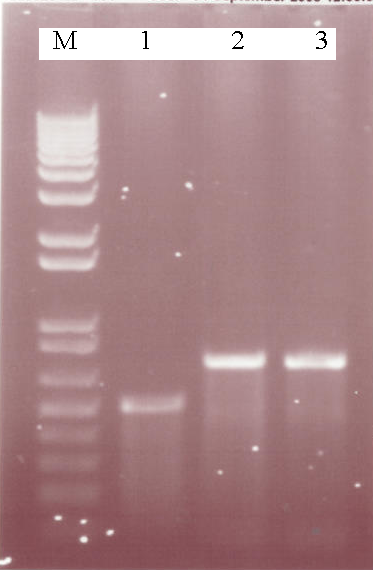
| + | [[Image:3-08PCR.PNG|thumb|200px|center|A 1% Agarose gel showing the results of various PCR reactions, M = Marker, Lane 1 - AmyE 5' Integration Sequence, Lane 2 - Aad9 PCR (58<sup>o</sup>C), Lane 3 - Aad9 PCR (56<sup>o</sup>C)]] | ||
| + | *The AmyE 5' integration sequence was succesfully produced by Pfu. This DNA can now be cut and ligated into a biobrick for use in our constructs. Aad9 was succesfully produced using Taq, providing conditions for use with Pfu for biobrick cloning | ||
| + | |||
| + | ==Dry Lab== | ||
| + | |||
| + | ===Motility=== | ||
| + | *Manual tracking of synthetic data was continued. Error bars were plotted and uploaded onto the OWW Wiki. | ||
| + | |||
| + | <br> | ||
| + | {{Imperial/EndPage|Notebook|Notebook}} | ||
Latest revision as of 20:44, 28 October 2008
3 September 2008WetlabCloning
Transformation of B.subtilis
ResultsUpper Lanes:
Lower Lanes:
Dry LabMotility
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"