Team:Paris/August 16
From 2008.igem.org
(Difference between revisions)
(→Construction of pLas-TetR-GFP tripart & rbs-LasR-dble ter) |
(→Digestion check from yesterday) |
||
| (3 intermediate revisions not shown) | |||
| Line 10: | Line 10: | ||
*10 µL of template DNA in 50 µL of pure water | *10 µL of template DNA in 50 µL of pure water | ||
*Blank : 10 µL of EB buffer in 50 µL of water. | *Blank : 10 µL of EB buffer in 50 µL of water. | ||
| - | {| | + | {||- style="text-align: center;" border="1" |
|align="center"|'''Digestion name''' | |align="center"|'''Digestion name''' | ||
|align="center"|'''What's in ?''' | |align="center"|'''What's in ?''' | ||
| - | |align="center"|'''DNA | + | |'''Enzymes''' |
| + | |align="center"|'''DNA C° (ng/µL)''' | ||
|- | |- | ||
|align="center"|D 158 | |align="center"|D 158 | ||
|align="center"| OmpR* | |align="center"| OmpR* | ||
| + | |XbaI-PstI | ||
|align="center"| 12 | |align="center"| 12 | ||
|- | |- | ||
|align="center"|D 159 | |align="center"|D 159 | ||
|align="center"| EnvZ* | |align="center"| EnvZ* | ||
| + | |XbaI-PstI | ||
|align="center"| 7 | |align="center"| 7 | ||
|- | |- | ||
|align="center"|D 102 | |align="center"|D 102 | ||
|align="center"|B0034 | |align="center"|B0034 | ||
| + | |SpeI-PstI | ||
|align="center"| 16 | |align="center"| 16 | ||
|} | |} | ||
| Line 59: | Line 63: | ||
L143, L144, T1 and T2 showed no colonies. The positive control with pUC19 worked well. | L143, L144, T1 and T2 showed no colonies. The positive control with pUC19 worked well. | ||
<br>We suppose that the ligation did not work. We will do it again today. | <br>We suppose that the ligation did not work. We will do it again today. | ||
| - | |||
==Cleaning of the DNA after the digestion== | ==Cleaning of the DNA after the digestion== | ||
We used the QIAcube to wash the DNA, following the [[Team:Paris/Notebook/Protocols#Purification_.28Kit_Promega.29|standard protocol.]] | We used the QIAcube to wash the DNA, following the [[Team:Paris/Notebook/Protocols#Purification_.28Kit_Promega.29|standard protocol.]] | ||
| - | |||
==Measure of DNA concentration of the digestion products== | ==Measure of DNA concentration of the digestion products== | ||
| Line 154: | Line 156: | ||
|4 | |4 | ||
|} | |} | ||
| - | |||
| - | |||
=Construction of pLas-TetR-GFP tripart & rbs-LasR-dble ter= | =Construction of pLas-TetR-GFP tripart & rbs-LasR-dble ter= | ||
| Line 202: | Line 202: | ||
|Description | |Description | ||
|Miniprep used | |Miniprep used | ||
| - | | | + | |Expected size |
| + | |Measured size | ||
|- | |- | ||
| | | | ||
| Line 214: | Line 215: | ||
|promoter J23101 - BV | |promoter J23101 - BV | ||
|1 | |1 | ||
| - | | | + | | |
| + | |style="background: #cbff7B"| | ||
|- | |- | ||
|D161 | |D161 | ||
| Line 220: | Line 222: | ||
|PTet (Tet promoter) - FI | |PTet (Tet promoter) - FI | ||
|1 | |1 | ||
| - | | | + | | |
| + | |style="background: #cbff7B"| | ||
|- | |- | ||
|D162 | |D162 | ||
| Line 226: | Line 229: | ||
|TetR - BI | |TetR - BI | ||
|1 | |1 | ||
| - | | | + | | |
| + | |style="background: #cbff7B"| | ||
|- | |- | ||
|D163 | |D163 | ||
| Line 232: | Line 236: | ||
|gfp generator - BV | |gfp generator - BV | ||
|2 | |2 | ||
| - | | | + | | |
| + | |style="background: #cbff7B"| | ||
|} | |} | ||
Latest revision as of 18:08, 4 September 2008
Construction of OmpR*+RBS and EnvZ*+RBS: LigationsCleaning of the DNA after the digestionWe used the QIAcube to wash the DNA, following the standard protocol. Measure of DNA concentration of the digestion productsWe used the biophotometer.
List of ligations
Creation of a registry of pFliL, pFlhDC, and FlhDCAnalysis of the transformation we did yesterdayL143, L144, T1 and T2 showed no colonies. The positive control with pUC19 worked well.
Cleaning of the DNA after the digestionWe used the QIAcube to wash the DNA, following the standard protocol. Measure of DNA concentration of the digestion productsWe used the biophotometer.
List of ligations
Construction of pLas-TetR-GFP tripart & rbs-LasR-dble terTransformation
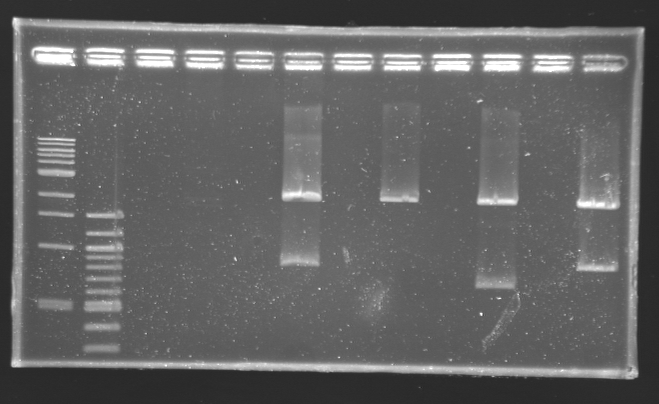
Digestion check from yesterday
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"